You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000002309_02622
You are here: Home > Sequence: MGYG000002309_02622
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Kurthia senegalensis | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Firmicutes; Bacilli; Bacillales_A; Planococcaceae; Kurthia; Kurthia senegalensis | |||||||||||
| CAZyme ID | MGYG000002309_02622 | |||||||||||
| CAZy Family | CBM50 | |||||||||||
| CAZyme Description | D-gamma-glutamyl-meso-diaminopimelic acid endopeptidase CwlS | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 21193; End: 22431 Strand: + | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| CBM50 | 162 | 203 | 3e-17 | 0.95 |
| CBM50 | 98 | 139 | 5.5e-17 | 0.95 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| PRK13914 | PRK13914 | 6.61e-43 | 12 | 412 | 8 | 481 | invasion associated endopeptidase. |
| PRK06347 | PRK06347 | 1.43e-33 | 29 | 268 | 328 | 591 | 1,4-beta-N-acetylmuramoylhydrolase. |
| pfam00877 | NLPC_P60 | 1.22e-28 | 307 | 410 | 1 | 104 | NlpC/P60 family. The function of this domain is unknown. It is found in several lipoproteins. |
| COG0791 | Spr | 7.18e-28 | 271 | 409 | 50 | 196 | Cell wall-associated hydrolase, NlpC family [Cell wall/membrane/envelope biogenesis]. |
| PRK06347 | PRK06347 | 6.90e-27 | 47 | 303 | 284 | 554 | 1,4-beta-N-acetylmuramoylhydrolase. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| QFF98463.1 | 2.42e-99 | 6 | 412 | 2 | 390 |
| VEI05004.1 | 4.41e-93 | 28 | 412 | 24 | 421 |
| VEI07037.1 | 8.21e-85 | 98 | 412 | 45 | 350 |
| QWU07155.1 | 7.62e-84 | 19 | 412 | 12 | 416 |
| AJD92285.1 | 1.44e-83 | 28 | 409 | 23 | 422 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 7CFL_A | 5.37e-15 | 298 | 407 | 17 | 132 | ChainA, Putative cell wall hydrolase phosphatase-associated protein [Clostridioides difficile],7CFL_B Chain B, Putative cell wall hydrolase phosphatase-associated protein [Clostridioides difficile],7CFL_C Chain C, Putative cell wall hydrolase phosphatase-associated protein [Clostridioides difficile],7CFL_D Chain D, Putative cell wall hydrolase phosphatase-associated protein [Clostridioides difficile] |
| 4XCM_A | 1.64e-12 | 293 | 408 | 112 | 226 | Crystalstructure of the putative NlpC/P60 D,L endopeptidase from T. thermophilus [Thermus thermophilus HB8],4XCM_B Crystal structure of the putative NlpC/P60 D,L endopeptidase from T. thermophilus [Thermus thermophilus HB8] |
| 4FDY_A | 8.36e-12 | 292 | 408 | 187 | 306 | ChainA, Similar to lipoprotein, NLP/P60 family [Staphylococcus aureus subsp. aureus Mu50],4FDY_B Chain B, Similar to lipoprotein, NLP/P60 family [Staphylococcus aureus subsp. aureus Mu50] |
| 6B8C_A | 5.05e-10 | 298 | 397 | 31 | 131 | Crystalstructure of NlpC/p60 domain of peptidoglycan hydrolase SagA [Enterococcus faecium] |
| 3PBC_A | 5.55e-08 | 294 | 394 | 84 | 194 | ChainA, Invasion Protein [Mycobacterium tuberculosis] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| O07532 | 9.27e-66 | 13 | 408 | 11 | 483 | Peptidoglycan endopeptidase LytF OS=Bacillus subtilis (strain 168) OX=224308 GN=lytF PE=1 SV=2 |
| O31852 | 1.23e-64 | 29 | 408 | 24 | 409 | D-gamma-glutamyl-meso-diaminopimelic acid endopeptidase CwlS OS=Bacillus subtilis (strain 168) OX=224308 GN=cwlS PE=1 SV=1 |
| P54421 | 1.82e-62 | 100 | 408 | 30 | 330 | Probable peptidoglycan endopeptidase LytE OS=Bacillus subtilis (strain 168) OX=224308 GN=lytE PE=1 SV=1 |
| Q49UX4 | 1.78e-46 | 11 | 207 | 4 | 197 | N-acetylmuramoyl-L-alanine amidase sle1 OS=Staphylococcus saprophyticus subsp. saprophyticus (strain ATCC 15305 / DSM 20229 / NCIMB 8711 / NCTC 7292 / S-41) OX=342451 GN=sle1 PE=3 SV=1 |
| Q01837 | 7.08e-44 | 96 | 412 | 196 | 524 | Probable endopeptidase p60 OS=Listeria ivanovii OX=1638 GN=iap PE=3 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as SP
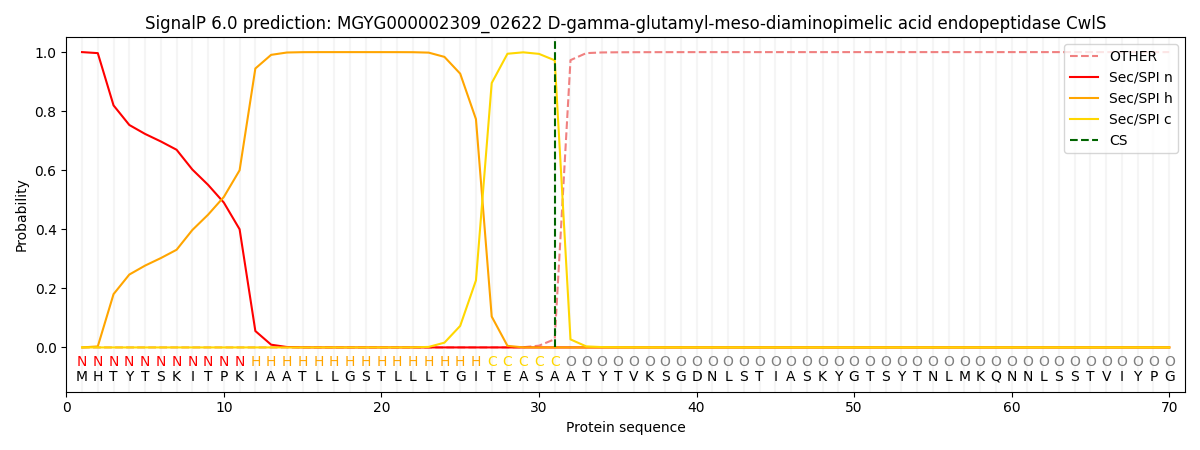
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 0.000246 | 0.999030 | 0.000179 | 0.000185 | 0.000166 | 0.000150 |
