You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000002310_00626
You are here: Home > Sequence: MGYG000002310_00626
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Microvirga massiliensis | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Proteobacteria; Alphaproteobacteria; Rhizobiales; Beijerinckiaceae; Microvirga; Microvirga massiliensis | |||||||||||
| CAZyme ID | MGYG000002310_00626 | |||||||||||
| CAZy Family | GH1 | |||||||||||
| CAZyme Description | hypothetical protein | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 640506; End: 642674 Strand: + | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH1 | 30 | 374 | 1.6e-42 | 0.7995337995337995 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| COG1091 | RfbD | 1.36e-64 | 442 | 707 | 2 | 279 | dTDP-4-dehydrorhamnose reductase [Cell wall/membrane/envelope biogenesis]. |
| cd05254 | dTDP_HR_like_SDR_e | 3.80e-61 | 442 | 664 | 1 | 230 | dTDP-6-deoxy-L-lyxo-4-hexulose reductase and related proteins, extended (e) SDRs. dTDP-6-deoxy-L-lyxo-4-hexulose reductase, an extended SDR, synthesizes dTDP-L-rhamnose from alpha-D-glucose-1-phosphate, providing the precursor of L-rhamnose, an essential cell wall component of many pathogenic bacteria. This subgroup has the characteristic active site tetrad and NADP-binding motif. This subgroup also contains human MAT2B, the regulatory subunit of methionine adenosyltransferase (MAT); MAT catalyzes S-adenosylmethionine synthesis. The human gene encoding MAT2B encodes two major splicing variants which are induced in human cell liver cancer and regulate HuR, an mRNA-binding protein which stabilizes the mRNA of several cyclins, to affect cell proliferation. Both MAT2B variants include this extended SDR domain. Extended SDRs are distinct from classical SDRs. In addition to the Rossmann fold (alpha/beta folding pattern with a central beta-sheet) core region typical of all SDRs, extended SDRs have a less conserved C-terminal extension of approximately 100 amino acids. Extended SDRs are a diverse collection of proteins, and include isomerases, epimerases, oxidoreductases, and lyases; they typically have a TGXXGXXG cofactor binding motif. SDRs are a functionally diverse family of oxidoreductases that have a single domain with a structurally conserved Rossmann fold, an NAD(P)(H)-binding region, and a structurally diverse C-terminal region. Sequence identity between different SDR enzymes is typically in the 15-30% range; they catalyze a wide range of activities including the metabolism of steroids, cofactors, carbohydrates, lipids, aromatic compounds, and amino acids, and act in redox sensing. Classical SDRs have an TGXXX[AG]XG cofactor binding motif and a YXXXK active site motif, with the Tyr residue of the active site motif serving as a critical catalytic residue (Tyr-151, human 15-hydroxyprostaglandin dehydrogenase numbering). In addition to the Tyr and Lys, there is often an upstream Ser and/or an Asn, contributing to the active site; while substrate binding is in the C-terminal region, which determines specificity. The standard reaction mechanism is a 4-pro-S hydride transfer and proton relay involving the conserved Tyr and Lys, a water molecule stabilized by Asn, and nicotinamide. Atypical SDRs generally lack the catalytic residues characteristic of the SDRs, and their glycine-rich NAD(P)-binding motif is often different from the forms normally seen in classical or extended SDRs. Complex (multidomain) SDRs such as ketoreductase domains of fatty acid synthase have a GGXGXXG NAD(P)-binding motif and an altered active site motif (YXXXN). Fungal type ketoacyl reductases have a TGXXXGX(1-2)G NAD(P)-binding motif. |
| pfam04321 | RmlD_sub_bind | 8.09e-56 | 443 | 707 | 1 | 282 | RmlD substrate binding domain. L-rhamnose is a saccharide required for the virulence of some bacteria. Its precursor, dTDP-L-rhamnose, is synthesized by four different enzymes the final one of which is RmlD. The RmlD substrate binding domain is responsible for binding a sugar nucleotide. |
| TIGR01214 | rmlD | 8.53e-56 | 442 | 707 | 1 | 286 | dTDP-4-dehydrorhamnose reductase. This enzyme catalyzes the last of 4 steps in making dTDP-rhamnose, a precursor of LPS core antigen, O-antigen, etc. [Cell envelope, Biosynthesis and degradation of surface polysaccharides and lipopolysaccharides] |
| PRK09987 | PRK09987 | 5.06e-20 | 476 | 722 | 39 | 292 | dTDP-4-dehydrorhamnose reductase; Provisional |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| QFU17850.1 | 0.0 | 2 | 713 | 3 | 715 |
| QRM34426.1 | 0.0 | 4 | 722 | 7 | 723 |
| QGM97233.1 | 2.71e-301 | 4 | 709 | 9 | 712 |
| QPQ56369.1 | 4.88e-291 | 4 | 709 | 5 | 707 |
| QIL01910.1 | 4.16e-287 | 4 | 704 | 12 | 712 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 3SC6_A | 1.57e-24 | 442 | 709 | 7 | 285 | 2.65Angstrom resolution crystal structure of dTDP-4-dehydrorhamnose reductase (rfbD) from Bacillus anthracis str. Ames in complex with NADP [Bacillus anthracis str. Ames],3SC6_B 2.65 Angstrom resolution crystal structure of dTDP-4-dehydrorhamnose reductase (rfbD) from Bacillus anthracis str. Ames in complex with NADP [Bacillus anthracis str. Ames],3SC6_C 2.65 Angstrom resolution crystal structure of dTDP-4-dehydrorhamnose reductase (rfbD) from Bacillus anthracis str. Ames in complex with NADP [Bacillus anthracis str. Ames],3SC6_D 2.65 Angstrom resolution crystal structure of dTDP-4-dehydrorhamnose reductase (rfbD) from Bacillus anthracis str. Ames in complex with NADP [Bacillus anthracis str. Ames],3SC6_E 2.65 Angstrom resolution crystal structure of dTDP-4-dehydrorhamnose reductase (rfbD) from Bacillus anthracis str. Ames in complex with NADP [Bacillus anthracis str. Ames],3SC6_F 2.65 Angstrom resolution crystal structure of dTDP-4-dehydrorhamnose reductase (rfbD) from Bacillus anthracis str. Ames in complex with NADP [Bacillus anthracis str. Ames] |
| 1KBZ_A | 6.73e-18 | 443 | 666 | 3 | 234 | ChainA, dTDP-glucose oxidoreductase [Salmonella enterica subsp. enterica serovar Typhimurium],1KC1_A Chain A, dTDP-glucose oxidoreductase [Salmonella enterica subsp. enterica serovar Typhimurium],1KC3_A Chain A, dTDP-glucose oxidoreductase [Salmonella enterica subsp. enterica serovar Typhimurium],1N2S_A Chain A, dTDP-glucose oxidoreductase [Salmonella enterica subsp. enterica serovar Typhimurium] |
| 6Z1H_A | 2.65e-11 | 30 | 365 | 63 | 412 | ChainA, ANCESTRAL RECONSTRUCTED GLYCOSIDASE [synthetic construct],6Z1H_B Chain B, ANCESTRAL RECONSTRUCTED GLYCOSIDASE [synthetic construct],6Z1M_A Chain A, Ancestral reconstructed glycosidase [synthetic construct],6Z1M_B Chain B, Ancestral reconstructed glycosidase [synthetic construct],6Z1M_C Chain C, Ancestral reconstructed glycosidase [synthetic construct] |
| 2YDX_A | 2.83e-11 | 441 | 682 | 3 | 259 | Crystalstructure of human S-adenosylmethionine synthetase 2, beta subunit [Homo sapiens],2YDX_B Crystal structure of human S-adenosylmethionine synthetase 2, beta subunit [Homo sapiens],2YDX_C Crystal structure of human S-adenosylmethionine synthetase 2, beta subunit [Homo sapiens],2YDX_D Crystal structure of human S-adenosylmethionine synthetase 2, beta subunit [Homo sapiens],2YDX_E Crystal structure of human S-adenosylmethionine synthetase 2, beta subunit [Homo sapiens] |
| 4NDN_E | 3.03e-11 | 441 | 682 | 18 | 274 | Structuralinsights of MAT enzymes: MATa2b complexed with SAM and PPNP [Homo sapiens],4NDN_F Structural insights of MAT enzymes: MATa2b complexed with SAM and PPNP [Homo sapiens] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| O66251 | 8.17e-30 | 442 | 670 | 3 | 238 | dTDP-4-dehydrorhamnose reductase OS=Aggregatibacter actinomycetemcomitans OX=714 GN=rmlD PE=1 SV=1 |
| P29781 | 7.28e-28 | 442 | 703 | 8 | 267 | dTDP-4-dehydrorhamnose reductase OS=Streptomyces griseus OX=1911 GN=strL PE=1 SV=1 |
| Q9L9E9 | 1.76e-27 | 442 | 659 | 4 | 222 | dTDP-4-keto-6-deoxy-D-glucose reductase OS=Streptomyces niveus OX=193462 GN=novS PE=3 SV=1 |
| A0QTF8 | 1.46e-25 | 415 | 673 | 12 | 267 | dTDP-4-dehydrorhamnose reductase OS=Mycolicibacterium smegmatis (strain ATCC 700084 / mc(2)155) OX=246196 GN=rmlD PE=1 SV=1 |
| P39631 | 2.07e-25 | 442 | 671 | 3 | 231 | Spore coat polysaccharide biosynthesis protein SpsK OS=Bacillus subtilis (strain 168) OX=224308 GN=spsK PE=3 SV=3 |
SignalP and Lipop Annotations help
This protein is predicted as OTHER
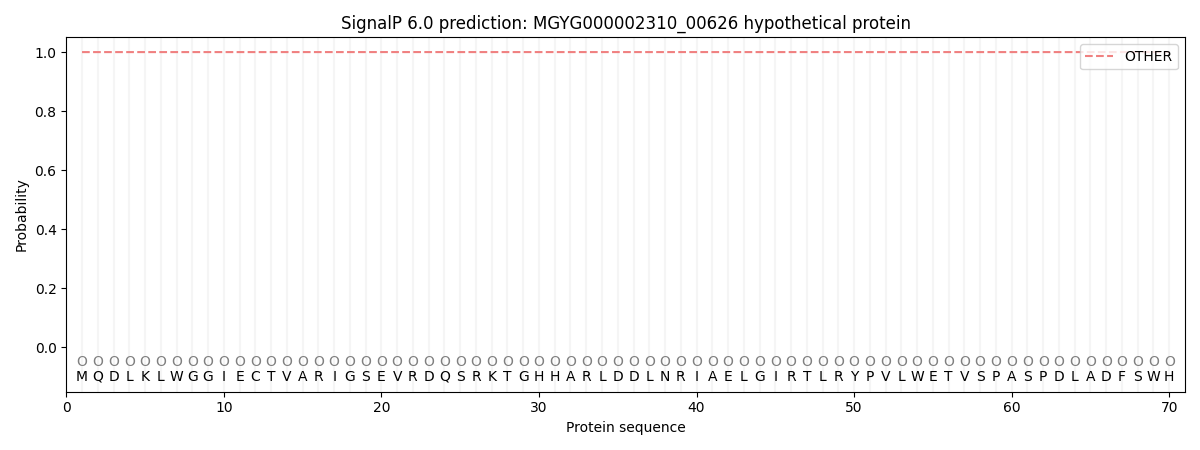
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 1.000046 | 0.000000 | 0.000000 | 0.000000 | 0.000000 | 0.000000 |
