You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000002363_02520
You are here: Home > Sequence: MGYG000002363_02520
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Raoultella ornithinolytica | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Proteobacteria; Gammaproteobacteria; Enterobacterales; Enterobacteriaceae; Raoultella; Raoultella ornithinolytica | |||||||||||
| CAZyme ID | MGYG000002363_02520 | |||||||||||
| CAZy Family | GH23 | |||||||||||
| CAZyme Description | Type IV secretion system protein virB1 | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 674839; End: 675474 Strand: - | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH23 | 15 | 153 | 2e-16 | 0.7925925925925926 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| cd16892 | LT_VirB1-like | 2.93e-76 | 7 | 154 | 1 | 143 | VirB1-like subfamily. This subfamily includes VirB1 protein, one of twelve proteins making up type IV secretion systems (T4SS). T4SS are macromolecular assemblies generally composed of VirB1-11 and VirD4 proteins, and are used by bacteria to transport material across their membranes. VirB1 acts as a lytic transglycosylase (LT), and is important with respect to piercing the peptidoglycan layer in the periplasm. LTs catalyze the cleavage of the beta-1,4-glycosidic bond between N-acetylmuramic acid (MurNAc) and N-acetyl-D-glucosamine (GlcNAc) as do "goose-type" lysozymes. However, in addition to this, they also make a new glycosidic bond with the C6 hydroxyl group of the same muramic acid residue. Proteins similar to this family include the soluble and insoluble membrane-bound LTs in bacteria, the LTs in bacteriophage lambda, as well as the eukaryotic "goose-type" lysozymes (goose egg-white lysozyme; GEWL). |
| PRK13864 | PRK13864 | 4.05e-13 | 7 | 135 | 33 | 161 | type IV secretion system lytic transglycosylase VirB1; Provisional |
| pfam01464 | SLT | 4.07e-07 | 6 | 104 | 1 | 76 | Transglycosylase SLT domain. This family is distantly related to pfam00062. Members are found in phages, type II, type III and type IV secretion systems. |
| cd16895 | TraH-like | 9.61e-06 | 2 | 107 | 1 | 109 | conjugal transfer protein H and similar proteins. This subfamily consists of several TraH proteins, putative conjugal transfer proteins of uncharacterized function, apparently found only in alphaproteobacteria. They have similarity to lysozyme and preserve the critical glutamate residue which has catalytic activity in lysozyme-like proteins. |
| PRK13843 | PRK13843 | 3.80e-05 | 1 | 107 | 1 | 110 | conjugal transfer protein TraH; Provisional |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| AXC30218.1 | 3.31e-150 | 1 | 211 | 1 | 211 |
| QWU12569.1 | 3.31e-150 | 1 | 211 | 1 | 211 |
| QLK16804.1 | 3.31e-150 | 1 | 211 | 1 | 211 |
| QLK21593.1 | 3.31e-150 | 1 | 211 | 1 | 211 |
| ATM21159.1 | 3.31e-150 | 1 | 211 | 1 | 211 |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| Q9RPY4 | 7.00e-55 | 1 | 189 | 1 | 186 | Type IV secretion system protein virB1 OS=Brucella suis biovar 1 (strain 1330) OX=204722 GN=virB1 PE=2 SV=1 |
| P0C398 | 2.81e-54 | 1 | 189 | 1 | 186 | Type IV secretion system protein virB1 OS=Brucella abortus biovar 1 (strain 9-941) OX=262698 GN=virB1 PE=3 SV=1 |
| Q2YIT5 | 2.81e-54 | 1 | 189 | 1 | 186 | Type IV secretion system protein virB1 OS=Brucella abortus (strain 2308) OX=359391 GN=virB1 PE=3 SV=1 |
| Q8YDZ5 | 2.81e-54 | 1 | 189 | 1 | 186 | Type IV secretion system protein virB1 OS=Brucella melitensis biotype 1 (strain 16M / ATCC 23456 / NCTC 10094) OX=224914 GN=virB1 PE=3 SV=1 |
| P0A3V6 | 2.77e-09 | 8 | 163 | 34 | 183 | Protein virB1 OS=Rhizobium radiobacter OX=358 GN=virB1 PE=3 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as OTHER
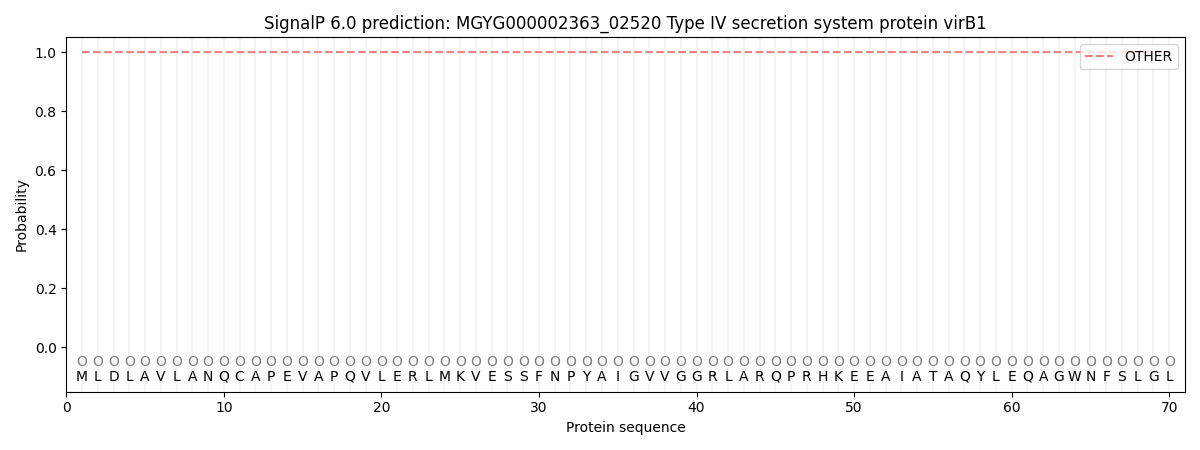
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 1.000039 | 0.000003 | 0.000000 | 0.000000 | 0.000000 | 0.000000 |
