You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000002372_01592
You are here: Home > Sequence: MGYG000002372_01592
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Clostridium_P perfringens | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Firmicutes_A; Clostridia; Clostridiales; Clostridiaceae; Clostridium_P; Clostridium_P perfringens | |||||||||||
| CAZyme ID | MGYG000002372_01592 | |||||||||||
| CAZy Family | GH84 | |||||||||||
| CAZyme Description | O-GlcNAcase NagJ | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 1735123; End: 1738128 Strand: - | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH84 | 182 | 479 | 1.3e-112 | 0.9932203389830508 |
| CBM32 | 632 | 762 | 1.7e-24 | 0.9516129032258065 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| pfam07555 | NAGidase | 1.01e-153 | 182 | 478 | 1 | 293 | beta-N-acetylglucosaminidase. This family has previously been described as a hyaluronidase. However, more recently it has been shown that this family has beta-N-acetylglucosaminidase activity. |
| pfam00754 | F5_F8_type_C | 1.74e-27 | 630 | 763 | 1 | 127 | F5/8 type C domain. This domain is also known as the discoidin (DS) domain family. |
| cd08546 | cohesin_like | 1.98e-23 | 777 | 908 | 1 | 144 | Cohesin domain, interaction parter of dockerin. Bacterial cohesin domains bind to a complementary protein domain named dockerin, and this interaction is required for the formation of the cellulosome, a cellulose-degrading complex. The cellulosome consists of scaffoldin, a noncatalytic scaffolding polypeptide, that comprises repeating cohesion modules and a single carbohydrate-binding module (CBM). Specific calcium-dependent interactions between cohesins and dockerins appear to be essential for cellulosome assembly. Cohesin modules are phylogenetically distributed into three groups: type I cohesin-dockerin interactions mediate assembly of a range of dockerin-borne enzymes to the complex, while type-II interactions mediate attachment of the cellulosome complex to the bacterial cell wall. Recently discovered type-III cohesins, such as found in the anchoring scaffoldin ScaE, appears to contribute to increased stability of the elaborate cellulosome complex. While the presence of cohesin and dockerin domains in a genome can be indicative of cellulolytic activity, cohesin domains may occur in a wider range of domain architectures, biological systems, and taxonomic lineages. |
| pfam02838 | Glyco_hydro_20b | 1.80e-22 | 44 | 175 | 2 | 123 | Glycosyl hydrolase family 20, domain 2. This domain has a zincin-like fold. |
| cd08523 | Reeler_cohesin_like | 5.51e-19 | 779 | 898 | 1 | 120 | Domains similar to the eukaryotic reeler domain and bacterial cohesins. This diverse family summarizes a set of distantly related domains, as revealed by structural similarity |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| AOY53993.1 | 0.0 | 1 | 1001 | 1 | 1001 |
| SQG39103.1 | 0.0 | 1 | 1001 | 1 | 1001 |
| AMN32783.1 | 0.0 | 1 | 1001 | 1 | 1001 |
| ABG84519.1 | 0.0 | 1 | 1001 | 1 | 1001 |
| ATD48631.1 | 0.0 | 1 | 1001 | 1 | 1001 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 2YDQ_A | 0.0 | 30 | 618 | 2 | 590 | CpOGAD298N in complex with hOGA-derived O-GlcNAc peptide [Clostridium perfringens],2YDR_A CpOGA D298N in complex with p53-derived O-GlcNAc peptide [Clostridium perfringens],2YDS_A CpOGA D298N in complex with TAB1-derived O-GlcNAc peptide [Clostridium perfringens] |
| 2V5D_A | 0.0 | 31 | 766 | 1 | 736 | Structureof a Family 84 Glycoside Hydrolase and a Family 32 Carbohydrate-Binding Module in Tandem from Clostridium perfringens. [Clostridium perfringens] |
| 2CBI_A | 0.0 | 31 | 624 | 1 | 594 | Structureof the Clostridium perfringens NagJ family 84 glycoside hydrolase, a homologue of human O-GlcNAcase [Clostridium perfringens],2CBI_B Structure of the Clostridium perfringens NagJ family 84 glycoside hydrolase, a homologue of human O-GlcNAcase [Clostridium perfringens],2CBJ_A Structure of the Clostridium perfringens NagJ family 84 glycoside hydrolase, a homologue of human O-GlcNAcase in complex with PUGNAc [Clostridium perfringens],2CBJ_B Structure of the Clostridium perfringens NagJ family 84 glycoside hydrolase, a homologue of human O-GlcNAcase in complex with PUGNAc [Clostridium perfringens],2V5C_A Family 84 glycoside hydrolase from Clostridium perfringens, 2.1 Angstrom structure [Clostridium perfringens],2V5C_B Family 84 glycoside hydrolase from Clostridium perfringens, 2.1 Angstrom structure [Clostridium perfringens],2VUR_A Chemical dissection of the link between Streptozotocin, O-GlcNAc and pancreatic cell death [Clostridium perfringens],2VUR_B Chemical dissection of the link between Streptozotocin, O-GlcNAc and pancreatic cell death [Clostridium perfringens],2X0Y_A Screening-based discovery of drug-like O-GlcNAcase inhibitor scaffolds [Clostridium perfringens],2X0Y_B Screening-based discovery of drug-like O-GlcNAcase inhibitor scaffolds [Clostridium perfringens] |
| 2XPK_A | 0.0 | 31 | 624 | 1 | 594 | Cell-penetrant,nanomolar O-GlcNAcase inhibitors selective against lysosomal hexosaminidases [Clostridium perfringens],2XPK_B Cell-penetrant, nanomolar O-GlcNAcase inhibitors selective against lysosomal hexosaminidases [Clostridium perfringens] |
| 7KHV_A | 0.0 | 31 | 624 | 1 | 594 | CpOGAIN COMPLEX WITH LIGAND 54 [Clostridium perfringens],7KHV_B CpOGA IN COMPLEX WITH LIGAND 54 [Clostridium perfringens],7KHV_C CpOGA IN COMPLEX WITH LIGAND 54 [Clostridium perfringens],7KHV_D CpOGA IN COMPLEX WITH LIGAND 54 [Clostridium perfringens],7KHV_E CpOGA IN COMPLEX WITH LIGAND 54 [Clostridium perfringens],7KHV_F CpOGA IN COMPLEX WITH LIGAND 54 [Clostridium perfringens] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| Q0TR53 | 0.0 | 1 | 1001 | 1 | 1001 | O-GlcNAcase NagJ OS=Clostridium perfringens (strain ATCC 13124 / DSM 756 / JCM 1290 / NCIMB 6125 / NCTC 8237 / Type A) OX=195103 GN=nagJ PE=1 SV=1 |
| Q8XL08 | 0.0 | 1 | 1001 | 1 | 1001 | O-GlcNAcase NagJ OS=Clostridium perfringens (strain 13 / Type A) OX=195102 GN=nagJ PE=1 SV=1 |
| Q89ZI2 | 3.08e-101 | 45 | 542 | 25 | 506 | O-GlcNAcase BT_4395 OS=Bacteroides thetaiotaomicron (strain ATCC 29148 / DSM 2079 / JCM 5827 / CCUG 10774 / NCTC 10582 / VPI-5482 / E50) OX=226186 GN=BT_4395 PE=1 SV=1 |
| P26831 | 1.70e-47 | 48 | 601 | 43 | 616 | Hyaluronoglucosaminidase OS=Clostridium perfringens (strain 13 / Type A) OX=195102 GN=nagH PE=1 SV=2 |
| O60502 | 3.12e-42 | 183 | 452 | 63 | 337 | Protein O-GlcNAcase OS=Homo sapiens OX=9606 GN=OGA PE=1 SV=2 |
SignalP and Lipop Annotations help
This protein is predicted as SP
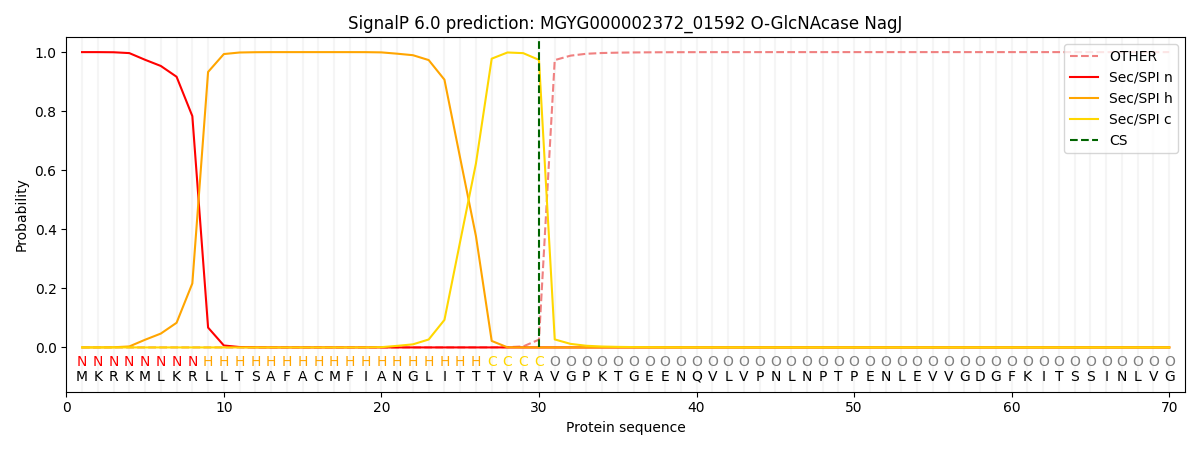
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 0.000309 | 0.998876 | 0.000231 | 0.000214 | 0.000182 | 0.000153 |
