You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000002381_02486
You are here: Home > Sequence: MGYG000002381_02486
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
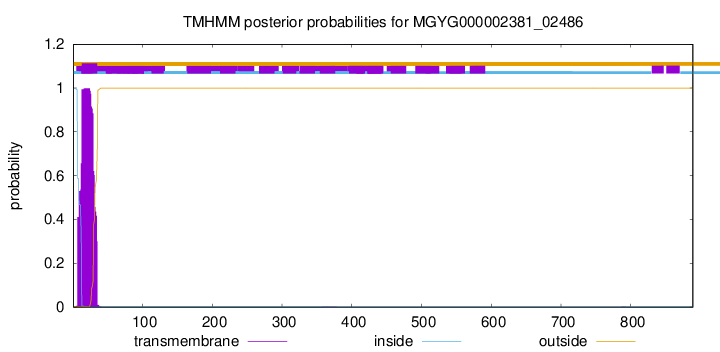
TMHMM annotations
Basic Information help
| Species | Stenotrophomonas bentonitica_A | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Proteobacteria; Gammaproteobacteria; Xanthomonadales; Xanthomonadaceae; Stenotrophomonas; Stenotrophomonas bentonitica_A | |||||||||||
| CAZyme ID | MGYG000002381_02486 | |||||||||||
| CAZy Family | GH2 | |||||||||||
| CAZyme Description | Exo-beta-D-glucosaminidase | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 76483; End: 79152 Strand: - | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH2 | 32 | 742 | 9.1e-110 | 0.6662234042553191 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| COG3250 | LacZ | 4.11e-119 | 36 | 889 | 18 | 808 | Beta-galactosidase/beta-glucuronidase [Carbohydrate transport and metabolism]. |
| pfam17786 | Mannosidase_ig | 1.89e-16 | 711 | 796 | 4 | 90 | Mannosidase Ig/CBM-like domain. This domain corresponds to domain 4 in the structure of Bacteroides thetaiotaomicron beta-mannosidase, BtMan2A. This domain has an Ig-like fold. |
| PRK10150 | PRK10150 | 8.51e-14 | 220 | 479 | 168 | 420 | beta-D-glucuronidase; Provisional |
| pfam17753 | Ig_mannosidase | 5.56e-13 | 824 | 885 | 15 | 78 | Ig-fold domain. This Ig-like fold domain is found in mannosidase enzymes. |
| pfam00703 | Glyco_hydro_2 | 2.05e-09 | 235 | 343 | 2 | 106 | Glycosyl hydrolases family 2. This family contains beta-galactosidase, beta-mannosidase and beta-glucuronidase activities. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| AOX63950.1 | 0.0 | 1 | 887 | 1 | 879 |
| AHY59190.1 | 0.0 | 1 | 888 | 1 | 880 |
| QHB71248.1 | 0.0 | 1 | 889 | 1 | 881 |
| QIO87992.1 | 0.0 | 25 | 885 | 17 | 871 |
| AOA72201.1 | 0.0 | 17 | 888 | 10 | 873 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 6BYE_A | 3.65e-318 | 37 | 888 | 12 | 855 | Crystalstructure of the GH2 exo-beta-mannanase from Xanthomonas axonopodis pv. citri in complex with mannose [Xanthomonas citri pv. citri str. 306],6BYE_B Crystal structure of the GH2 exo-beta-mannanase from Xanthomonas axonopodis pv. citri in complex with mannose [Xanthomonas citri pv. citri str. 306] |
| 6BYC_A | 3.86e-318 | 37 | 888 | 12 | 855 | Crystalstructure of the GH2 exo-beta-mannanase from Xanthomonas axonopodis pv. citri [Xanthomonas citri pv. citri str. 306] |
| 6BYI_A | 2.78e-317 | 37 | 888 | 10 | 853 | Crystalstructure of the acid-base mutant (E477A) of the GH2 exo-beta-mannanase from Xanthomonas axonopodis pv. citri [Xanthomonas citri pv. citri str. 306],6BYI_B Crystal structure of the acid-base mutant (E477A) of the GH2 exo-beta-mannanase from Xanthomonas axonopodis pv. citri [Xanthomonas citri pv. citri str. 306] |
| 6BYG_A | 2.88e-317 | 37 | 888 | 12 | 855 | Crystalstructure of the nucleophile mutant (E575A) of the GH2 exo-beta-mannanase from Xanthomonas axonopodis pv. citri [Xanthomonas citri pv. citri str. 306],6BYG_B Crystal structure of the nucleophile mutant (E575A) of the GH2 exo-beta-mannanase from Xanthomonas axonopodis pv. citri [Xanthomonas citri pv. citri str. 306] |
| 2VJX_A | 2.19e-156 | 53 | 867 | 21 | 818 | Structuraland biochemical evidence for a boat-like transition state in beta-mannosidases [Bacteroides thetaiotaomicron VPI-5482],2VJX_B Structural and biochemical evidence for a boat-like transition state in beta-mannosidases [Bacteroides thetaiotaomicron VPI-5482],2VL4_A Structural and biochemical evidence for a boat-like transition state in beta-mannosidases [Bacteroides thetaiotaomicron VPI-5482],2VL4_B Structural and biochemical evidence for a boat-like transition state in beta-mannosidases [Bacteroides thetaiotaomicron VPI-5482],2VMF_A Structural and biochemical evidence for a boat-like transition state in beta-mannosidases [Bacteroides thetaiotaomicron VPI-5482],2VMF_B Structural and biochemical evidence for a boat-like transition state in beta-mannosidases [Bacteroides thetaiotaomicron VPI-5482],2VO5_A Structural and biochemical evidence for a boat-like transition state in beta-mannosidases [Bacteroides thetaiotaomicron VPI-5482],2VO5_B Structural and biochemical evidence for a boat-like transition state in beta-mannosidases [Bacteroides thetaiotaomicron VPI-5482],2VOT_A Structural and biochemical evidence for a boat-like transition state in beta-mannosidases [Bacteroides thetaiotaomicron VPI-5482],2VOT_B Structural and biochemical evidence for a boat-like transition state in beta-mannosidases [Bacteroides thetaiotaomicron VPI-5482],2VQT_A Structural and biochemical evidence for a boat-like transition state in beta-mannosidases [Bacteroides thetaiotaomicron],2VQT_B Structural and biochemical evidence for a boat-like transition state in beta-mannosidases [Bacteroides thetaiotaomicron],2VR4_A Transition-state mimicry in mannoside hydrolysis: characterisation of twenty six inhibitors and insight into binding from linear free energy relationships and 3-D structure [Bacteroides thetaiotaomicron VPI-5482],2VR4_B Transition-state mimicry in mannoside hydrolysis: characterisation of twenty six inhibitors and insight into binding from linear free energy relationships and 3-D structure [Bacteroides thetaiotaomicron VPI-5482] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| I2C092 | 4.10e-110 | 32 | 863 | 8 | 836 | Beta-mannosidase B OS=Thermothelomyces thermophilus OX=78579 GN=man9 PE=1 SV=1 |
| Q95327 | 2.84e-105 | 59 | 886 | 41 | 878 | Beta-mannosidase OS=Capra hircus OX=9925 GN=MANBA PE=1 SV=1 |
| Q29444 | 3.95e-105 | 59 | 886 | 41 | 878 | Beta-mannosidase OS=Bos taurus OX=9913 GN=MANBA PE=1 SV=1 |
| Q4WAH4 | 2.74e-102 | 32 | 866 | 8 | 829 | Beta-mannosidase B OS=Neosartorya fumigata (strain ATCC MYA-4609 / Af293 / CBS 101355 / FGSC A1100) OX=330879 GN=mndB PE=3 SV=1 |
| Q8K2I4 | 2.92e-102 | 59 | 886 | 41 | 878 | Beta-mannosidase OS=Mus musculus OX=10090 GN=Manba PE=1 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as SP
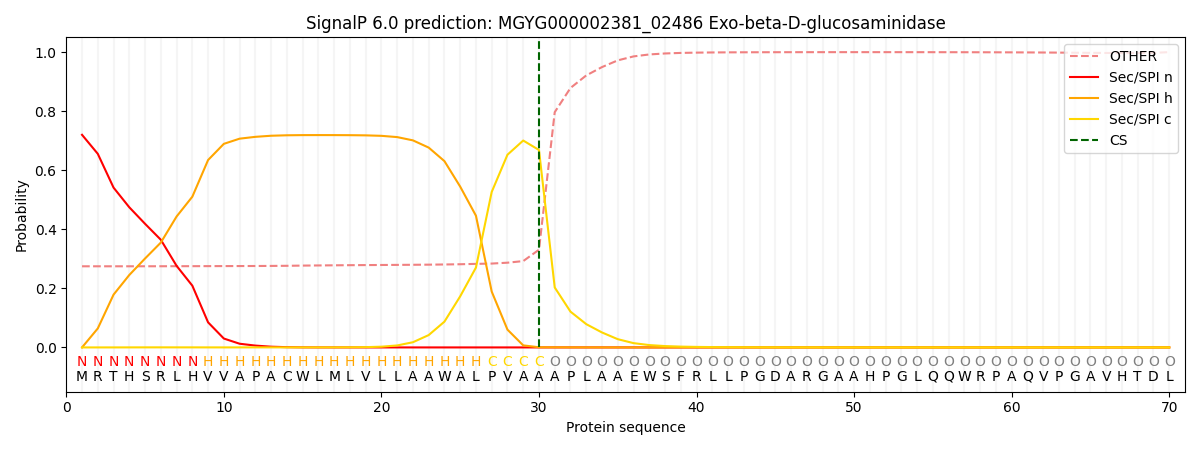
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 0.297230 | 0.693637 | 0.006345 | 0.000899 | 0.000556 | 0.001310 |