You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000002383_02602
You are here: Home > Sequence: MGYG000002383_02602
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
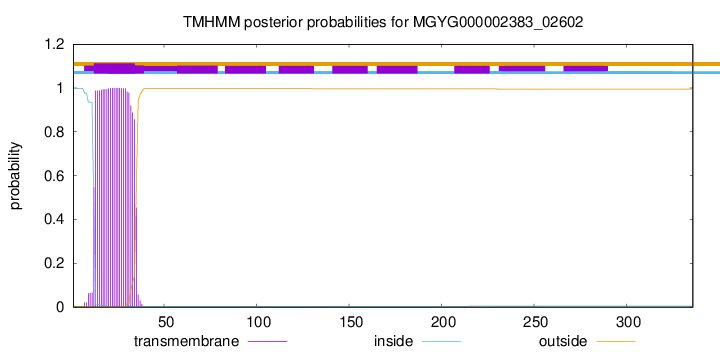
TMHMM annotations
Basic Information help
| Species | Lacticaseibacillus zeae | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Firmicutes; Bacilli; Lactobacillales; Lactobacillaceae; Lacticaseibacillus; Lacticaseibacillus zeae | |||||||||||
| CAZyme ID | MGYG000002383_02602 | |||||||||||
| CAZy Family | GH73 | |||||||||||
| CAZyme Description | hypothetical protein | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 11718; End: 12728 Strand: - | |||||||||||
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| COG0791 | Spr | 1.03e-18 | 162 | 324 | 17 | 181 | Cell wall-associated hydrolase, NlpC family [Cell wall/membrane/envelope biogenesis]. |
| pfam00877 | NLPC_P60 | 1.96e-17 | 227 | 314 | 1 | 81 | NlpC/P60 family. The function of this domain is unknown. It is found in several lipoproteins. |
| PRK13914 | PRK13914 | 2.22e-16 | 172 | 311 | 321 | 454 | invasion associated endopeptidase. |
| NF033741 | NlpC_p60_RipA | 1.46e-15 | 218 | 324 | 331 | 441 | NlpC/P60 family peptidoglycan endopeptidase RipA. |
| NF033742 | NlpC_p60_RipB | 3.05e-11 | 195 | 312 | 54 | 178 | NlpC/P60 family peptidoglycan endopeptidase RipB. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| ARE43723.1 | 6.45e-240 | 1 | 336 | 1 | 336 |
| QOP54548.1 | 9.16e-240 | 1 | 336 | 1 | 336 |
| QBA75848.1 | 3.73e-239 | 1 | 336 | 1 | 336 |
| QTH67767.1 | 3.06e-238 | 1 | 336 | 1 | 336 |
| QKK92482.1 | 3.06e-238 | 1 | 336 | 1 | 336 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 6B8C_A | 1.70e-28 | 210 | 334 | 24 | 140 | Crystalstructure of NlpC/p60 domain of peptidoglycan hydrolase SagA [Enterococcus faecium] |
| 3NE0_A | 2.72e-11 | 139 | 326 | 34 | 200 | Structureand functional regulation of RipA, a mycobacterial enzyme essential for daughter cell separation [Mycobacterium tuberculosis H37Rv] |
| 3PBC_A | 2.72e-11 | 139 | 326 | 34 | 200 | ChainA, Invasion Protein [Mycobacterium tuberculosis] |
| 7CFL_A | 4.43e-11 | 218 | 311 | 17 | 107 | ChainA, Putative cell wall hydrolase phosphatase-associated protein [Clostridioides difficile],7CFL_B Chain B, Putative cell wall hydrolase phosphatase-associated protein [Clostridioides difficile],7CFL_C Chain C, Putative cell wall hydrolase phosphatase-associated protein [Clostridioides difficile],7CFL_D Chain D, Putative cell wall hydrolase phosphatase-associated protein [Clostridioides difficile] |
| 3S0Q_A | 9.40e-11 | 139 | 326 | 35 | 201 | ChainA, INVASION PROTEIN [Mycobacterium tuberculosis] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| P13692 | 1.56e-25 | 210 | 334 | 398 | 514 | Protein P54 OS=Enterococcus faecium OX=1352 PE=3 SV=2 |
| Q01835 | 9.06e-13 | 206 | 312 | 387 | 485 | Probable endopeptidase p60 OS=Listeria grayi OX=1641 GN=iap PE=3 SV=1 |
| Q736M3 | 3.17e-11 | 218 | 324 | 214 | 315 | Gamma-D-glutamyl-L-lysine dipeptidyl-peptidase OS=Bacillus cereus (strain ATCC 10987 / NRS 248) OX=222523 GN=ykfC PE=1 SV=1 |
| P96740 | 7.91e-11 | 218 | 328 | 169 | 275 | Gamma-DL-glutamyl hydrolase OS=Bacillus subtilis (strain 168) OX=224308 GN=pgdS PE=1 SV=2 |
| Q01838 | 1.29e-10 | 218 | 311 | 411 | 496 | Probable endopeptidase p60 OS=Listeria seeligeri OX=1640 GN=iap PE=3 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as OTHER

| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 1.000021 | 0.000013 | 0.000000 | 0.000000 | 0.000000 | 0.000004 |