You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000002407_00580
You are here: Home > Sequence: MGYG000002407_00580
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Cohnella sp900169535 | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Firmicutes; Bacilli; Paenibacillales; Paenibacillaceae; Cohnella; Cohnella sp900169535 | |||||||||||
| CAZyme ID | MGYG000002407_00580 | |||||||||||
| CAZy Family | CBM8 | |||||||||||
| CAZyme Description | hypothetical protein | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 104670; End: 108083 Strand: - | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH44 | 632 | 1135 | 1.7e-137 | 0.9902723735408561 |
| CBM8 | 474 | 614 | 5.2e-33 | 0.9790209790209791 |
| CBM8 | 236 | 375 | 7e-26 | 0.972027972027972 |
| CBM8 | 42 | 177 | 4.5e-23 | 0.965034965034965 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| pfam12891 | Glyco_hydro_44 | 8.25e-90 | 688 | 922 | 2 | 234 | Glycoside hydrolase family 44. This is a family of bacterial glycoside hydrolases formerly known as cellulase family J, and now known as Cel44A. It is one of the major enzymatic components of the cellulosome of Clostridium thermocellum strain F1 and of many other Firmicutes. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| QNK59691.1 | 0.0 | 14 | 1134 | 14 | 1128 |
| QTH43477.1 | 0.0 | 7 | 1134 | 8 | 1126 |
| AYQ73506.1 | 1.21e-310 | 37 | 1134 | 83 | 1240 |
| QJD86673.1 | 7.65e-297 | 12 | 1134 | 13 | 1195 |
| AVP98921.1 | 4.90e-149 | 471 | 1137 | 26 | 719 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 3FW6_A | 1.27e-104 | 631 | 1134 | 18 | 531 | Crystalstructure of CelM2, a bifunctional glucanase-xylanase protein from a metagenome library [uncultured bacterium] |
| 3II1_A | 1.30e-104 | 631 | 1134 | 18 | 531 | Structuralcharacterization of difunctional glucanase-xylanse CelM2 [uncultured bacterium] |
| 2E0P_A | 1.13e-53 | 633 | 1077 | 10 | 454 | ChainA, Endoglucanase [Acetivibrio thermocellus],2E4T_A Chain A, Endoglucanase [Acetivibrio thermocellus],2EO7_A Chain A, Endoglucanase [Acetivibrio thermocellus] |
| 2EEX_A | 2.79e-53 | 633 | 1077 | 10 | 454 | ChainA, Endoglucanase [Acetivibrio thermocellus],2EJ1_A Chain A, Endoglucanase [Acetivibrio thermocellus],2EQD_A Chain A, Endoglucanase [Acetivibrio thermocellus] |
| 3IK2_A | 9.19e-47 | 633 | 1104 | 3 | 478 | CrystalStructure of a Glycoside Hydrolase Family 44 Endoglucanase produced by Clostridium acetobutylium ATCC 824 [Clostridium acetobutylicum ATCC 824] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| P22533 | 5.05e-46 | 633 | 1126 | 788 | 1294 | Beta-mannanase/endoglucanase A OS=Caldicellulosiruptor saccharolyticus OX=44001 GN=manA PE=1 SV=2 |
| P29719 | 7.41e-45 | 633 | 1077 | 36 | 481 | Endoglucanase A OS=Paenibacillus lautus OX=1401 GN=celA PE=3 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as SP
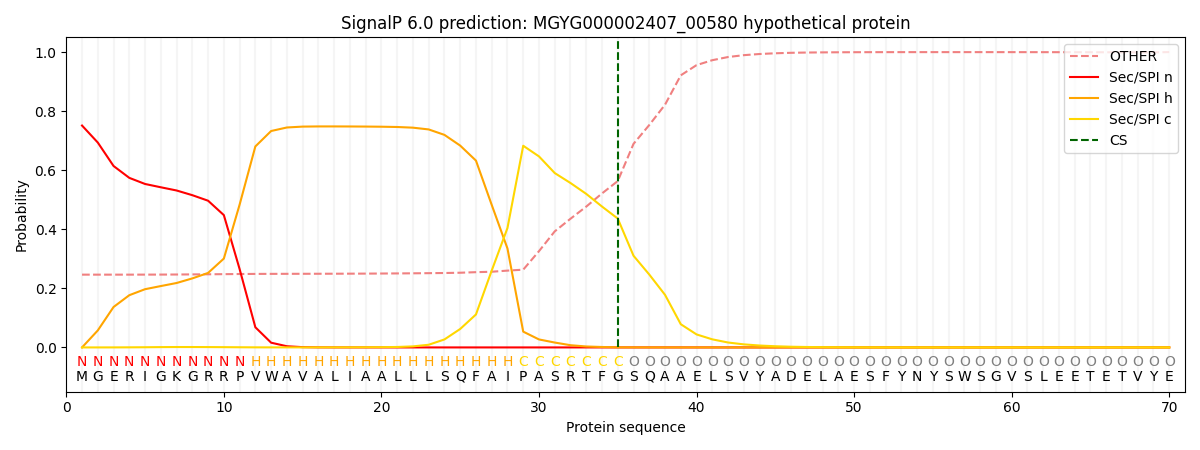
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 0.253392 | 0.742742 | 0.002840 | 0.000393 | 0.000272 | 0.000328 |