You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000002408_00044
You are here: Home > Sequence: MGYG000002408_00044
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Bacillus_BD tuaregi | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Firmicutes; Bacilli; Bacillales_B; DSM-18226; Bacillus_BD; Bacillus_BD tuaregi | |||||||||||
| CAZyme ID | MGYG000002408_00044 | |||||||||||
| CAZy Family | CBM50 | |||||||||||
| CAZyme Description | hypothetical protein | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 9583; End: 10569 Strand: - | |||||||||||
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| cd06583 | PGRP | 1.02e-29 | 22 | 166 | 2 | 126 | Peptidoglycan recognition proteins (PGRPs) are pattern recognition receptors that bind, and in certain cases, hydrolyze peptidoglycans (PGNs) of bacterial cell walls. PGRPs have been divided into three classes: short PGRPs (PGRP-S), that are small (20 kDa) extracellular proteins; intermediate PGRPs (PGRP-I) that are 40-45 kDa and are predicted to be transmembrane proteins; and long PGRPs (PGRP-L), up to 90 kDa, which may be either intracellular or transmembrane. Several structures of PGRPs are known in insects and mammals, some bound with substrates like Muramyl Tripeptide (MTP) or Tracheal Cytotoxin (TCT). The substrate binding site is conserved in PGRP-LCx, PGRP-LE, and PGRP-Ialpha proteins. This family includes Zn-dependent N-Acetylmuramoyl-L-alanine Amidase, EC:3.5.1.28. This enzyme cleaves the amide bond between N-acetylmuramoyl and L-amino acids, preferentially D-lactyl-L-Ala, in bacterial cell walls. The structure for the bacteriophage T7 lysozyme shows that two of the conserved histidines and a cysteine are zinc binding residues. Site-directed mutagenesis of T7 lysozyme indicates that two conserved residues, a Tyr and a Lys, are important for amidase activity. |
| pfam01510 | Amidase_2 | 3.05e-20 | 22 | 154 | 2 | 116 | N-acetylmuramoyl-L-alanine amidase. This family includes zinc amidases that have N-acetylmuramoyl-L-alanine amidase activity EC:3.5.1.28. This enzyme domain cleaves the amide bond between N-acetylmuramoyl and L-amino acids in bacterial cell walls (preferentially: D-lactyl-L-Ala). The structure is known for the bacteriophage T7 structure and shows that two of the conserved histidines are zinc binding. |
| smart00644 | Ami_2 | 2.78e-17 | 22 | 154 | 3 | 123 | Ami_2 domain. |
| COG5632 | CwlA | 9.87e-09 | 21 | 154 | 23 | 146 | N-acetylmuramoyl-L-alanine amidase CwlA [Cell wall/membrane/envelope biogenesis]. |
| pfam05036 | SPOR | 6.61e-06 | 292 | 327 | 1 | 36 | Sporulation related domain. This 70 residue domain is composed of two 35 residue repeats found in proteins involved in sporulation and cell division such as FtsN, DedD, and CwlM. This domain is involved in binding peptidoglycan. Two tandem repeats fold into a pseudo-2-fold symmetric single-domain structure containing numerous contacts between the repeats. FtsN is an essential cell division protein with a simple bitopic topology, a short N-terminal cytoplasmic segment fused to a large carboxy periplasmic domain through a single transmembrane domain. These repeats lay at the periplasmic C-terminus. FtsN localizes to the septum ring complex. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| AFV12491.1 | 8.30e-155 | 1 | 301 | 1 | 293 |
| AST58407.1 | 2.37e-154 | 1 | 301 | 1 | 293 |
| CUH93073.1 | 6.78e-154 | 1 | 301 | 1 | 293 |
| ANX01801.1 | 1.94e-153 | 1 | 301 | 1 | 293 |
| AQS59659.1 | 2.25e-152 | 1 | 301 | 1 | 293 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 3HMB_A | 1.60e-10 | 17 | 124 | 21 | 136 | ChainA, N-acetylmuramoyl-L-alanine amidase xlyA [Bacillus subtilis],3HMB_B Chain B, N-acetylmuramoyl-L-alanine amidase xlyA [Bacillus subtilis],3HMB_C Chain C, N-acetylmuramoyl-L-alanine amidase xlyA [Bacillus subtilis] |
| 3RDR_A | 6.62e-09 | 17 | 124 | 21 | 136 | Structureof the catalytic domain of XlyA [Bacillus subtilis] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| Q99125 | 1.13e-09 | 1 | 229 | 2 | 227 | Probable N-acetylmuramoyl-L-alanine amidase OS=Bacillus licheniformis OX=1402 PE=3 SV=1 |
| P36550 | 1.60e-08 | 1 | 233 | 2 | 232 | N-acetylmuramoyl-L-alanine amidase CwlL OS=Bacillus licheniformis OX=1402 GN=cwlL PE=3 SV=1 |
| P39800 | 7.01e-08 | 17 | 188 | 18 | 160 | N-acetylmuramoyl-L-alanine amidase XlyA OS=Bacillus subtilis (strain 168) OX=224308 GN=xlyA PE=1 SV=1 |
| P54450 | 2.38e-06 | 17 | 166 | 18 | 147 | N-acetylmuramoyl-L-alanine amidase CwlH OS=Bacillus subtilis (strain 168) OX=224308 GN=cwlH PE=1 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as OTHER
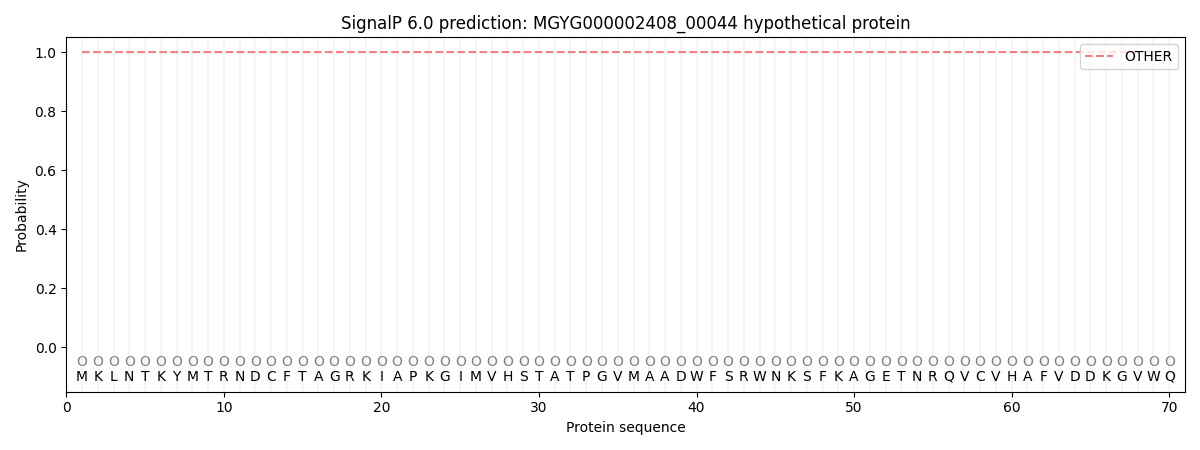
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 1.000036 | 0.000000 | 0.000000 | 0.000000 | 0.000000 | 0.000000 |
