You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000002424_00051
You are here: Home > Sequence: MGYG000002424_00051
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
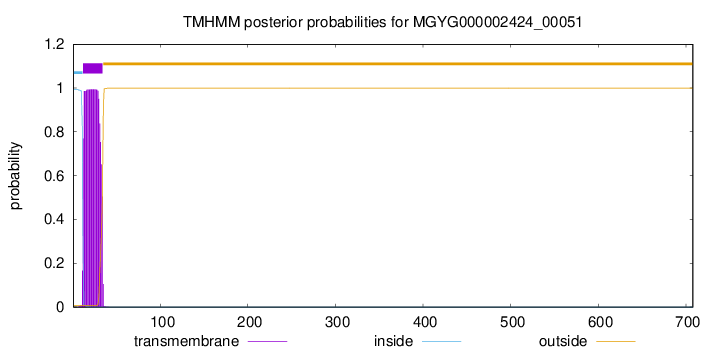
TMHMM annotations
Basic Information help
| Species | Monoglobus pectinilyticus | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Firmicutes_A; Clostridia; Monoglobales; Monoglobaceae; Monoglobus; Monoglobus pectinilyticus | |||||||||||
| CAZyme ID | MGYG000002424_00051 | |||||||||||
| CAZy Family | PL1 | |||||||||||
| CAZyme Description | hypothetical protein | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 52686; End: 54812 Strand: + | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| PL1 | 230 | 431 | 6.2e-29 | 0.8960396039603961 |
| CBM13 | 563 | 707 | 2.1e-19 | 0.6968085106382979 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| COG3866 | PelB | 1.63e-22 | 169 | 515 | 17 | 343 | Pectate lyase [Carbohydrate transport and metabolism]. |
| pfam14200 | RicinB_lectin_2 | 2.16e-18 | 560 | 643 | 10 | 89 | Ricin-type beta-trefoil lectin domain-like. |
| smart00656 | Amb_all | 2.51e-15 | 245 | 431 | 12 | 187 | Amb_all domain. |
| pfam14200 | RicinB_lectin_2 | 1.10e-13 | 603 | 692 | 5 | 89 | Ricin-type beta-trefoil lectin domain-like. |
| NF035929 | lectin_1 | 9.47e-10 | 570 | 702 | 718 | 834 | lectin. Lectins are important adhesin proteins, which bind carbohydrate structures on host cell surface. The carbohydrate specificity of diverse lectins to a large extent dictates bacteria tissue tropism by mediating specific attachment to unique host sites expressing the corresponding carbohydrate receptor. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| AUO18238.1 | 0.0 | 1 | 708 | 1 | 708 |
| CDM70399.1 | 1.20e-291 | 2 | 708 | 1 | 697 |
| ASR46637.1 | 6.92e-165 | 3 | 532 | 5 | 536 |
| ANY68265.1 | 6.90e-157 | 2 | 532 | 4 | 538 |
| CBL17651.1 | 5.31e-101 | 15 | 702 | 10 | 685 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 3KRG_A | 5.21e-07 | 279 | 428 | 149 | 323 | ChainA, Pectate lyase [Bacillus subtilis] |
| 5AMV_A | 1.60e-06 | 279 | 428 | 149 | 323 | Structuralinsights into the loss of catalytic competence in pectate lyase at low pH [Bacillus subtilis],5X2I_A Polygalacturonate Lyase by Fusing with a Self-assembling Amphipathic Peptide [Bacillus subtilis subsp. subtilis str. 168] |
| 1BN8_A | 1.68e-06 | 279 | 428 | 170 | 344 | BacillusSubtilis Pectate Lyase [Bacillus subtilis] |
| 3ZSC_A | 1.75e-06 | 199 | 400 | 20 | 213 | Catalyticfunction and substrate recognition of the pectate lyase from Thermotoga maritima [Thermotoga maritima] |
| 2BSP_A | 3.87e-06 | 279 | 428 | 170 | 344 | ChainA, PROTEIN (PECTATE LYASE) [Bacillus subtilis] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| P94449 | 3.71e-17 | 181 | 513 | 34 | 336 | Pectin lyase OS=Bacillus subtilis OX=1423 GN=pelB PE=1 SV=1 |
| O34819 | 4.98e-17 | 181 | 494 | 34 | 320 | Pectin lyase OS=Bacillus subtilis (strain 168) OX=224308 GN=pelB PE=3 SV=1 |
| P27027 | 1.06e-16 | 182 | 504 | 10 | 297 | Pectin lyase OS=Pseudomonas marginalis OX=298 GN=pnl PE=1 SV=2 |
| Q00645 | 5.95e-15 | 196 | 486 | 43 | 319 | Pectate lyase plyB OS=Emericella nidulans (strain FGSC A4 / ATCC 38163 / CBS 112.46 / NRRL 194 / M139) OX=227321 GN=plyB PE=1 SV=1 |
| B8NBC2 | 3.48e-14 | 196 | 471 | 43 | 303 | Probable pectate lyase B OS=Aspergillus flavus (strain ATCC 200026 / FGSC A1120 / IAM 13836 / NRRL 3357 / JCM 12722 / SRRC 167) OX=332952 GN=plyB PE=3 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as SP

| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 0.000480 | 0.998803 | 0.000221 | 0.000150 | 0.000147 | 0.000143 |