You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000002424_00914
You are here: Home > Sequence: MGYG000002424_00914
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
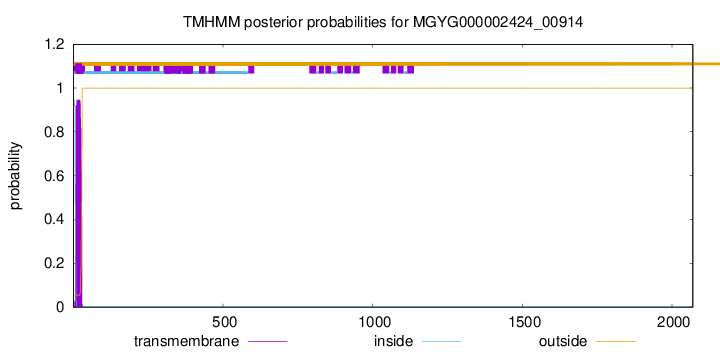
TMHMM annotations
Basic Information help
| Species | Monoglobus pectinilyticus | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Firmicutes_A; Clostridia; Monoglobales; Monoglobaceae; Monoglobus; Monoglobus pectinilyticus | |||||||||||
| CAZyme ID | MGYG000002424_00914 | |||||||||||
| CAZy Family | PL1 | |||||||||||
| CAZyme Description | hypothetical protein | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 1028615; End: 1034827 Strand: - | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| PL1 | 725 | 919 | 6.2e-42 | 0.9836065573770492 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| smart00656 | Amb_all | 2.81e-10 | 745 | 891 | 16 | 161 | Amb_all domain. |
| pfam00395 | SLH | 1.90e-09 | 1947 | 1988 | 1 | 42 | S-layer homology domain. |
| COG3866 | PelB | 9.02e-08 | 664 | 924 | 14 | 276 | Pectate lyase [Carbohydrate transport and metabolism]. |
| pfam00395 | SLH | 1.03e-07 | 1886 | 1927 | 1 | 42 | S-layer homology domain. |
| NF033190 | inl_like_NEAT_1 | 8.66e-07 | 1824 | 1996 | 507 | 691 | NEAT domain-containing leucine-rich repeat protein. Members of this family have an N-terminal NEAT (near transporter) domain often associated with iron transport, followed by a leucine-rich repeat region with significant sequence similarity to the internalins of Listeria monocytogenes. However, since Bacillus cereus (from which this protein was described, in PMID:16978259) is not considered an intracellular pathogen, and the function may be iron transport rather than internalization, applying the name "internalin" to this family probably would be misleading. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| AUO19083.1 | 0.0 | 1 | 2070 | 1 | 2070 |
| QIH34580.1 | 1.35e-45 | 664 | 934 | 19 | 275 |
| AUO19049.1 | 6.97e-41 | 666 | 2062 | 40 | 1227 |
| AWW32652.1 | 1.32e-40 | 674 | 942 | 47 | 301 |
| APZ48036.1 | 2.71e-40 | 674 | 920 | 26 | 263 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 6BT4_A | 2.74e-11 | 1884 | 2067 | 24 | 201 | Crystalstructure of the SLH domain of Sap from Bacillus anthracis in complex with a pyruvylated SCWP unit [Bacillus anthracis] |
| 3PYW_A | 2.80e-11 | 1884 | 2067 | 3 | 180 | Thestructure of the SLH domain from B. anthracis surface array protein at 1.8A [Bacillus anthracis] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| B8NQQ7 | 4.44e-33 | 671 | 999 | 19 | 303 | Probable pectate lyase C OS=Aspergillus flavus (strain ATCC 200026 / FGSC A1120 / IAM 13836 / NRRL 3357 / JCM 12722 / SRRC 167) OX=332952 GN=plyC PE=3 SV=1 |
| Q0CLG7 | 2.67e-32 | 671 | 953 | 19 | 272 | Probable pectate lyase C OS=Aspergillus terreus (strain NIH 2624 / FGSC A1156) OX=341663 GN=plyC PE=3 SV=1 |
| Q2UB83 | 2.91e-31 | 671 | 999 | 19 | 303 | Probable pectate lyase C OS=Aspergillus oryzae (strain ATCC 42149 / RIB 40) OX=510516 GN=plyC PE=3 SV=1 |
| A1DPF0 | 5.83e-30 | 671 | 1050 | 20 | 351 | Probable pectate lyase C OS=Neosartorya fischeri (strain ATCC 1020 / DSM 3700 / CBS 544.65 / FGSC A1164 / JCM 1740 / NRRL 181 / WB 181) OX=331117 GN=plyC PE=3 SV=1 |
| B0XMA2 | 1.42e-29 | 671 | 958 | 20 | 278 | Probable pectate lyase C OS=Neosartorya fumigata (strain CEA10 / CBS 144.89 / FGSC A1163) OX=451804 GN=plyC PE=3 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as SP
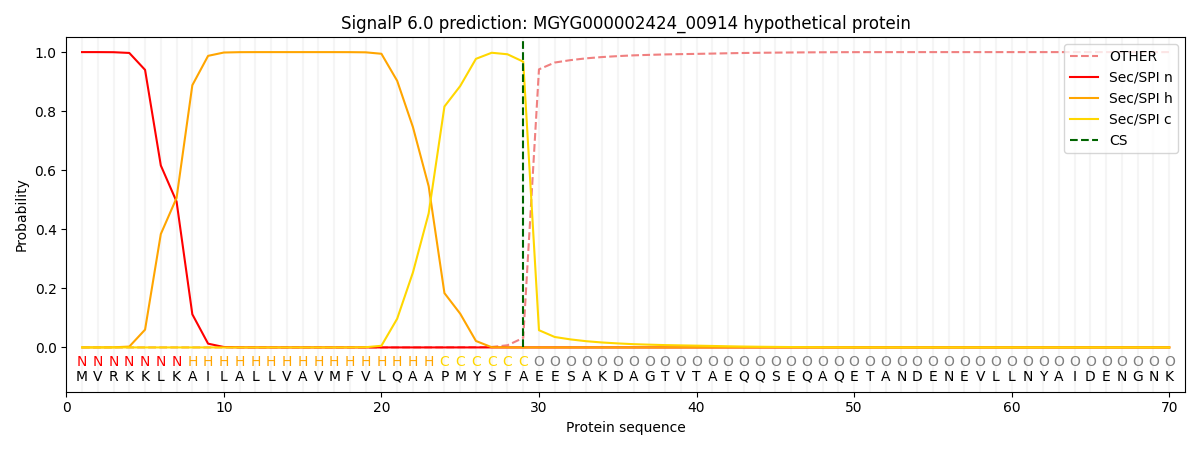
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 0.000309 | 0.998992 | 0.000191 | 0.000182 | 0.000157 | 0.000148 |