You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000002455_02312
You are here: Home > Sequence: MGYG000002455_02312
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Bacteroides cellulosilyticus | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Bacteroidota; Bacteroidia; Bacteroidales; Bacteroidaceae; Bacteroides; Bacteroides cellulosilyticus | |||||||||||
| CAZyme ID | MGYG000002455_02312 | |||||||||||
| CAZy Family | GH30 | |||||||||||
| CAZyme Description | hypothetical protein | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 97867; End: 99474 Strand: - | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH30 | 42 | 531 | 1.6e-179 | 0.993801652892562 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| pfam14587 | Glyco_hydr_30_2 | 3.34e-130 | 39 | 390 | 1 | 357 | O-Glycosyl hydrolase family 30. |
| COG5520 | XynC | 7.77e-17 | 1 | 534 | 1 | 430 | O-Glycosyl hydrolase [Cell wall/membrane/envelope biogenesis]. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| ALJ59966.1 | 0.0 | 1 | 535 | 1 | 535 |
| QUT89021.1 | 0.0 | 1 | 535 | 1 | 535 |
| BBK87114.1 | 3.28e-293 | 36 | 535 | 24 | 520 |
| QUT97738.1 | 4.04e-293 | 36 | 535 | 39 | 535 |
| QUT34917.1 | 5.73e-293 | 36 | 535 | 39 | 535 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 3CLW_A | 1.25e-72 | 39 | 531 | 5 | 495 | Crystalstructure of conserved exported protein from Bacteroides fragilis [Bacteroides fragilis NCTC 9343],3CLW_B Crystal structure of conserved exported protein from Bacteroides fragilis [Bacteroides fragilis NCTC 9343],3CLW_C Crystal structure of conserved exported protein from Bacteroides fragilis [Bacteroides fragilis NCTC 9343],3CLW_D Crystal structure of conserved exported protein from Bacteroides fragilis [Bacteroides fragilis NCTC 9343],3CLW_E Crystal structure of conserved exported protein from Bacteroides fragilis [Bacteroides fragilis NCTC 9343],3CLW_F Crystal structure of conserved exported protein from Bacteroides fragilis [Bacteroides fragilis NCTC 9343] |
| 4FMV_A | 1.74e-14 | 42 | 533 | 5 | 387 | CrystalStructure Analysis of a GH30 Endoxylanase from Clostridium papyrosolvens C71 [Ruminiclostridium papyrosolvens DSM 2782] |
| 7O0E_A | 2.24e-13 | 42 | 531 | 5 | 446 | ChainA, GH30 family xylanase [Thermothelomyces thermophilus ATCC 42464],7O0E_G Chain G, GH30 family xylanase [Thermothelomyces thermophilus ATCC 42464] |
| 7NCX_AAA | 5.76e-13 | 42 | 531 | 12 | 453 | ChainAAA, GH30 family xylanase [Thermothelomyces thermophilus ATCC 42464] |
| 6M5Z_A | 7.03e-13 | 42 | 485 | 25 | 408 | ChainA, GH30 Xylanase C [Talaromyces cellulolyticus CF-2612],6M5Z_B Chain B, GH30 Xylanase C [Talaromyces cellulolyticus CF-2612] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| Q76FP5 | 8.27e-16 | 42 | 445 | 24 | 383 | Endo-beta-1,6-galactanase OS=Hypocrea rufa OX=5547 GN=6GAL PE=1 SV=1 |
| G2Q1N4 | 2.43e-13 | 42 | 531 | 29 | 470 | GH30 family xylanase OS=Myceliophthora thermophila (strain ATCC 42464 / BCRC 31852 / DSM 1799) OX=573729 GN=Xyn30A PE=1 SV=1 |
| Q45070 | 3.51e-06 | 42 | 533 | 36 | 421 | Glucuronoxylanase XynC OS=Bacillus subtilis (strain 168) OX=224308 GN=xynC PE=1 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as LIPO
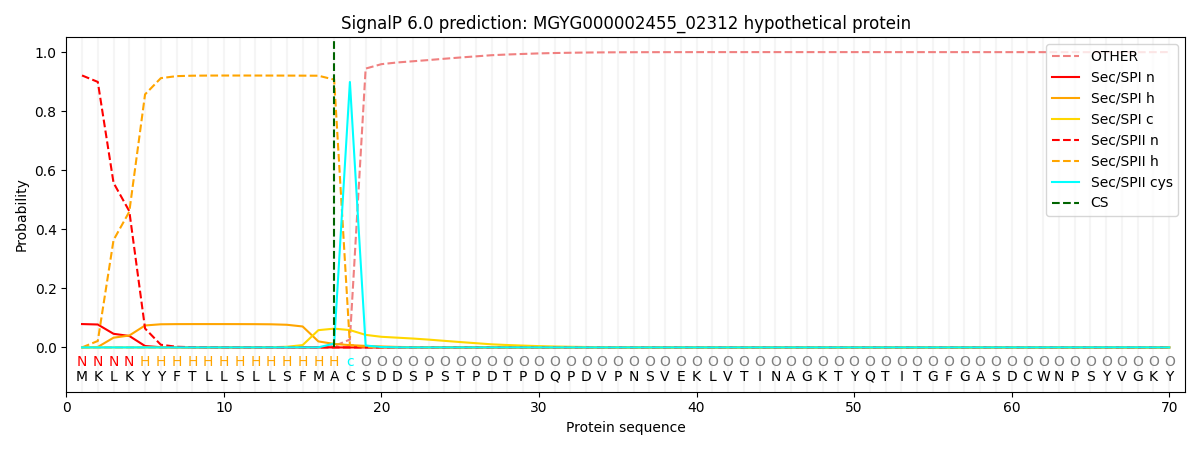
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 0.000083 | 0.077589 | 0.922233 | 0.000034 | 0.000037 | 0.000031 |
