You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000002468_01153
You are here: Home > Sequence: MGYG000002468_01153
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Enterococcus caccae | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Firmicutes; Bacilli; Lactobacillales; Enterococcaceae; Enterococcus; Enterococcus caccae | |||||||||||
| CAZyme ID | MGYG000002468_01153 | |||||||||||
| CAZy Family | GH39 | |||||||||||
| CAZyme Description | HTH-type transcriptional activator RhaS | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 171178; End: 173535 Strand: + | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH39 | 333 | 732 | 1.5e-34 | 0.962877030162413 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| COG2207 | AraC | 3.49e-29 | 156 | 270 | 9 | 123 | AraC-type DNA-binding domain and AraC-containing proteins [Transcription]. |
| smart00342 | HTH_ARAC | 8.49e-29 | 183 | 266 | 1 | 84 | helix_turn_helix, arabinose operon control protein. |
| COG4977 | GlxA | 3.02e-26 | 156 | 267 | 209 | 320 | Transcriptional regulator GlxA family, contains an amidase domain and an AraC-type DNA-binding HTH domain [Transcription]. |
| pfam12833 | HTH_18 | 1.44e-25 | 189 | 268 | 1 | 81 | Helix-turn-helix domain. |
| COG2169 | AdaA | 1.46e-23 | 165 | 267 | 80 | 180 | Methylphosphotriester-DNA--protein-cysteine methyltransferase (N-terminal fragment of Ada), contains Zn-binding and two AraC-type DNA-binding domains [Replication, recombination and repair]. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| QHZ53252.1 | 5.54e-170 | 4 | 784 | 2 | 775 |
| QNN74361.1 | 2.55e-100 | 8 | 778 | 6 | 760 |
| AVF22981.1 | 5.69e-99 | 333 | 784 | 108 | 549 |
| QYF80459.1 | 1.55e-96 | 9 | 784 | 10 | 771 |
| AQR76238.1 | 4.40e-96 | 349 | 784 | 5 | 430 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 1PX8_A | 1.54e-17 | 369 | 779 | 62 | 477 | Crystalstructure of beta-D-xylosidase from Thermoanaerobacterium saccharolyticum, a family 39 glycoside hydrolase [Thermoanaerobacterium saccharolyticum],1PX8_B Crystal structure of beta-D-xylosidase from Thermoanaerobacterium saccharolyticum, a family 39 glycoside hydrolase [Thermoanaerobacterium saccharolyticum],1UHV_A Crystal structure of beta-D-xylosidase from Thermoanaerobacterium saccharolyticum, a family 39 glycoside hydrolase [Thermoanaerobacterium saccharolyticum],1UHV_B Crystal structure of beta-D-xylosidase from Thermoanaerobacterium saccharolyticum, a family 39 glycoside hydrolase [Thermoanaerobacterium saccharolyticum],1UHV_C Crystal structure of beta-D-xylosidase from Thermoanaerobacterium saccharolyticum, a family 39 glycoside hydrolase [Thermoanaerobacterium saccharolyticum],1UHV_D Crystal structure of beta-D-xylosidase from Thermoanaerobacterium saccharolyticum, a family 39 glycoside hydrolase [Thermoanaerobacterium saccharolyticum] |
| 2BFG_A | 1.08e-15 | 321 | 778 | 6 | 477 | crystalstructure of beta-xylosidase (fam GH39) in complex with dinitrophenyl-beta-xyloside and covalently bound xyloside [Geobacillus stearothermophilus],2BFG_B crystal structure of beta-xylosidase (fam GH39) in complex with dinitrophenyl-beta-xyloside and covalently bound xyloside [Geobacillus stearothermophilus],2BFG_C crystal structure of beta-xylosidase (fam GH39) in complex with dinitrophenyl-beta-xyloside and covalently bound xyloside [Geobacillus stearothermophilus],2BFG_D crystal structure of beta-xylosidase (fam GH39) in complex with dinitrophenyl-beta-xyloside and covalently bound xyloside [Geobacillus stearothermophilus],2BFG_E crystal structure of beta-xylosidase (fam GH39) in complex with dinitrophenyl-beta-xyloside and covalently bound xyloside [Geobacillus stearothermophilus],2BFG_F crystal structure of beta-xylosidase (fam GH39) in complex with dinitrophenyl-beta-xyloside and covalently bound xyloside [Geobacillus stearothermophilus],2BFG_G crystal structure of beta-xylosidase (fam GH39) in complex with dinitrophenyl-beta-xyloside and covalently bound xyloside [Geobacillus stearothermophilus],2BFG_H crystal structure of beta-xylosidase (fam GH39) in complex with dinitrophenyl-beta-xyloside and covalently bound xyloside [Geobacillus stearothermophilus] |
| 2BS9_A | 1.89e-15 | 321 | 778 | 6 | 477 | Nativecrystal structure of a GH39 beta-xylosidase XynB1 from Geobacillus stearothermophilus [Geobacillus stearothermophilus],2BS9_B Native crystal structure of a GH39 beta-xylosidase XynB1 from Geobacillus stearothermophilus [Geobacillus stearothermophilus],2BS9_C Native crystal structure of a GH39 beta-xylosidase XynB1 from Geobacillus stearothermophilus [Geobacillus stearothermophilus],2BS9_D Native crystal structure of a GH39 beta-xylosidase XynB1 from Geobacillus stearothermophilus [Geobacillus stearothermophilus],2BS9_E Native crystal structure of a GH39 beta-xylosidase XynB1 from Geobacillus stearothermophilus [Geobacillus stearothermophilus],2BS9_F Native crystal structure of a GH39 beta-xylosidase XynB1 from Geobacillus stearothermophilus [Geobacillus stearothermophilus],2BS9_G Native crystal structure of a GH39 beta-xylosidase XynB1 from Geobacillus stearothermophilus [Geobacillus stearothermophilus],2BS9_H Native crystal structure of a GH39 beta-xylosidase XynB1 from Geobacillus stearothermophilus [Geobacillus stearothermophilus] |
| 1W91_A | 1.02e-14 | 321 | 778 | 6 | 477 | crystalstructure of 1,4-BETA-D-XYLAN XYLOHYDROLASE solve using anomalous signal from Seleniomethionine [synthetic construct],1W91_B crystal structure of 1,4-BETA-D-XYLAN XYLOHYDROLASE solve using anomalous signal from Seleniomethionine [synthetic construct],1W91_C crystal structure of 1,4-BETA-D-XYLAN XYLOHYDROLASE solve using anomalous signal from Seleniomethionine [synthetic construct],1W91_D crystal structure of 1,4-BETA-D-XYLAN XYLOHYDROLASE solve using anomalous signal from Seleniomethionine [synthetic construct],1W91_E crystal structure of 1,4-BETA-D-XYLAN XYLOHYDROLASE solve using anomalous signal from Seleniomethionine [synthetic construct],1W91_F crystal structure of 1,4-BETA-D-XYLAN XYLOHYDROLASE solve using anomalous signal from Seleniomethionine [synthetic construct],1W91_G crystal structure of 1,4-BETA-D-XYLAN XYLOHYDROLASE solve using anomalous signal from Seleniomethionine [synthetic construct],1W91_H crystal structure of 1,4-BETA-D-XYLAN XYLOHYDROLASE solve using anomalous signal from Seleniomethionine [synthetic construct] |
| 6UQJ_A | 1.71e-13 | 342 | 713 | 41 | 433 | Crystalstructure of the GH39 enzyme from Xanthomonas axonopodis pv. citri [Xanthomonas citri pv. citri str. 306] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| Q5HJR8 | 5.55e-43 | 27 | 770 | 23 | 730 | Uncharacterized HTH-type transcriptional regulator SACOL0084 OS=Staphylococcus aureus (strain COL) OX=93062 GN=SACOL0084 PE=4 SV=2 |
| Q99XB1 | 1.00e-42 | 27 | 770 | 23 | 730 | Uncharacterized HTH-type transcriptional regulator SAV0101 OS=Staphylococcus aureus (strain Mu50 / ATCC 700699) OX=158878 GN=SAV0101 PE=4 SV=1 |
| Q7A882 | 1.00e-42 | 27 | 770 | 23 | 730 | Uncharacterized HTH-type transcriptional regulator SA0097 OS=Staphylococcus aureus (strain N315) OX=158879 GN=SA0097 PE=4 SV=1 |
| Q6GD21 | 2.42e-42 | 27 | 770 | 23 | 730 | Uncharacterized HTH-type transcriptional regulator SAS0078 OS=Staphylococcus aureus (strain MSSA476) OX=282459 GN=SAS0078 PE=4 SV=1 |
| Q8NYT6 | 2.42e-42 | 27 | 770 | 23 | 730 | Uncharacterized HTH-type transcriptional regulator MW0077 OS=Staphylococcus aureus (strain MW2) OX=196620 GN=MW0077 PE=4 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as OTHER
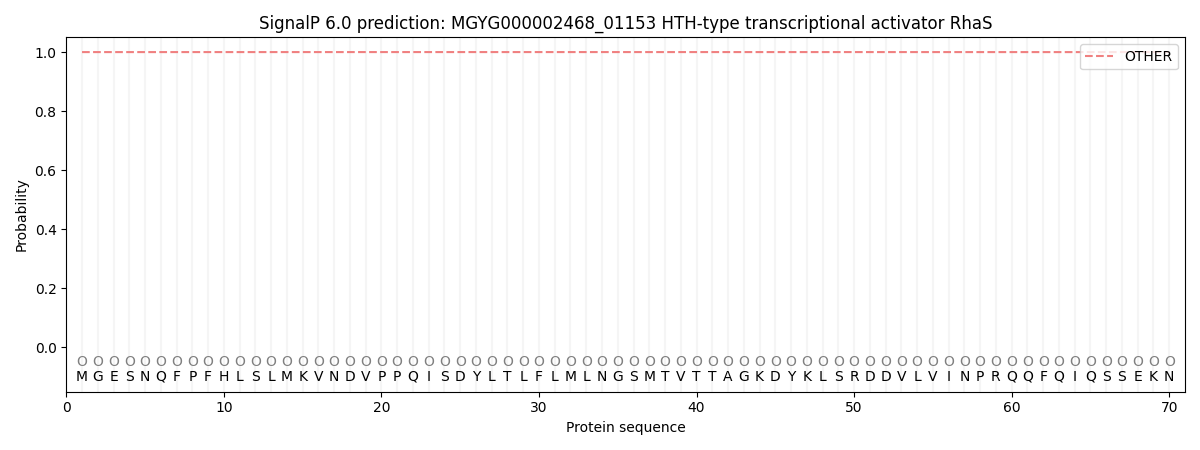
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 1.000065 | 0.000000 | 0.000000 | 0.000000 | 0.000000 | 0.000000 |
