You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000002473_00934
You are here: Home > Sequence: MGYG000002473_00934
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
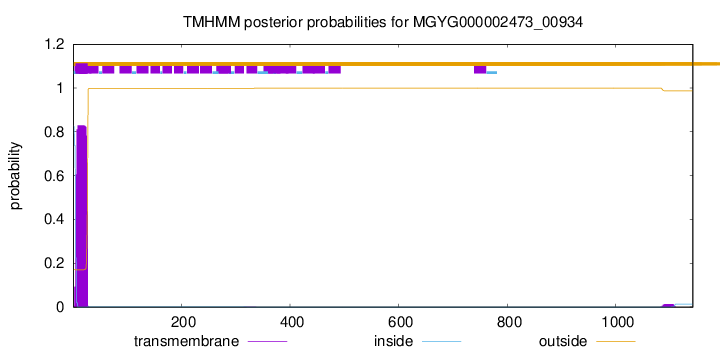
TMHMM annotations
Basic Information help
| Species | Victivallis vadensis | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Verrucomicrobiota; Lentisphaeria; Victivallales; Victivallaceae; Victivallis; Victivallis vadensis | |||||||||||
| CAZyme ID | MGYG000002473_00934 | |||||||||||
| CAZy Family | CBM67 | |||||||||||
| CAZyme Description | hypothetical protein | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 20049; End: 23477 Strand: + | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| CBM67 | 23 | 163 | 5.1e-33 | 0.9829545454545454 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| cd09621 | CBM9_like_5 | 1.62e-57 | 960 | 1139 | 1 | 187 | DOMON-like type 9 carbohydrate binding module. Family 9 carbohydrate-binding modules (CBM9) play a role in the microbial degradation of cellulose and hemicellulose (materials found in plants). The domain has previously been called cellulose-binding domain. The polysaccharide binding sites of CBMs with available 3D structure have been found to be either flat surfaces with interactions formed by predominantly aromatic residues (tryptophan and tyrosine), or extended shallow grooves. CBM9 domains found in this uncharacterized heterogeneous subfamily are often located at the C-terminus of longer proteins and may co-occur with various other functional domains such as glycosyl hydrolases. The CBM9 module in these architectures may be involved in binding to carbohydrates. |
| pfam08531 | Bac_rhamnosid_N | 1.04e-11 | 46 | 155 | 1 | 153 | Alpha-L-rhamnosidase N-terminal domain. This family consists of bacterial rhamnosidase A and B enzymes. This domain is probably involved in substrate recognition. |
| cd00241 | DOMON_like | 6.96e-07 | 975 | 1122 | 5 | 158 | Domon-like ligand-binding domains. DOMON-like domains can be found in all three kindgoms of life and are a diverse group of ligand binding domains that have been shown to interact with sugars and hemes. DOMON domains were initially thought to confer protein-protein interactions. They were subsequently found as a heme-binding motif in cellobiose dehydrogenase, an extracellular fungal oxidoreductase that degrades both lignin and cellulose, and in ethylbenzene dehydrogenase, an enzyme that aids in the anaerobic degradation of hydrocarbons. The domain interacts with sugars in the type 9 carbohydrate binding modules (CBM9), which are present in a variety of glycosyl hydrolases, and it can also be found at the N-terminus of sensor histidine kinases. |
| cd09619 | CBM9_like_4 | 0.002 | 982 | 1137 | 30 | 180 | DOMON-like type 9 carbohydrate binding module. Family 9 carbohydrate-binding modules (CBM9) play a role in the microbial degradation of cellulose and hemicellulose (materials found in plants). The domain has previously been called cellulose-binding domain. The polysaccharide binding sites of CBMs with available 3D structure have been found to be either flat surfaces with interactions formed by predominantly aromatic residues (tryptophan and tyrosine), or extended shallow grooves. CBM9 domains found in this uncharacterized heterogeneous subfamily are often located at the C-terminus of longer proteins and may co-occur with various other domains. |
| PRK10340 | ebgA | 0.002 | 27 | 98 | 102 | 176 | cryptic beta-D-galactosidase subunit alpha; Reviewed |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| AVM46501.1 | 0.0 | 5 | 1140 | 11 | 1153 |
| AHF92778.1 | 4.44e-304 | 11 | 1140 | 27 | 1177 |
| AVM46714.1 | 6.10e-196 | 190 | 1140 | 26 | 968 |
| AVM46271.1 | 2.95e-168 | 190 | 1141 | 22 | 966 |
| AVM46712.1 | 2.21e-162 | 172 | 1140 | 60 | 1011 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 6T0Q_A | 6.99e-12 | 23 | 163 | 200 | 344 | PleurotusOstreatus Lectin (POL), apo form [Pleurotus ostreatus] |
| 6T1D_A | 7.16e-12 | 23 | 163 | 203 | 347 | PleurotusOstreatus Lectin (POL), compelx with melibiose [Pleurotus ostreatus],6T1D_B Pleurotus Ostreatus Lectin (POL), compelx with melibiose [Pleurotus ostreatus],6T1D_C Pleurotus Ostreatus Lectin (POL), compelx with melibiose [Pleurotus ostreatus],6T1D_D Pleurotus Ostreatus Lectin (POL), compelx with melibiose [Pleurotus ostreatus],6T1D_E Pleurotus Ostreatus Lectin (POL), compelx with melibiose [Pleurotus ostreatus],6T1D_F Pleurotus Ostreatus Lectin (POL), compelx with melibiose [Pleurotus ostreatus] |
| 6KBJ_A | 8.09e-12 | 23 | 163 | 220 | 364 | Structureof Lectin from Pleurotus ostreatus in complex with malonate [Pleurotus ostreatus],6KBQ_A Crystal Structure of Lectin from Pleurotus ostreatus in complex with Glycerol [Pleurotus ostreatus],6KC2_A Crystal Structure of Lectin from Pleurotus ostreatus in complex with Rhamnose [Pleurotus ostreatus],6LI7_A Crystal Structure of Lectin from Pleurotus ostreatus in complex with GalNAc [Pleurotus ostreatus],6LIK_A Crystal Structure of Lectin from Pleurotus ostreatus in complex with Galactose [Pleurotus ostreatus] |
Swiss-Prot Hits help
SignalP and Lipop Annotations help
This protein is predicted as SP
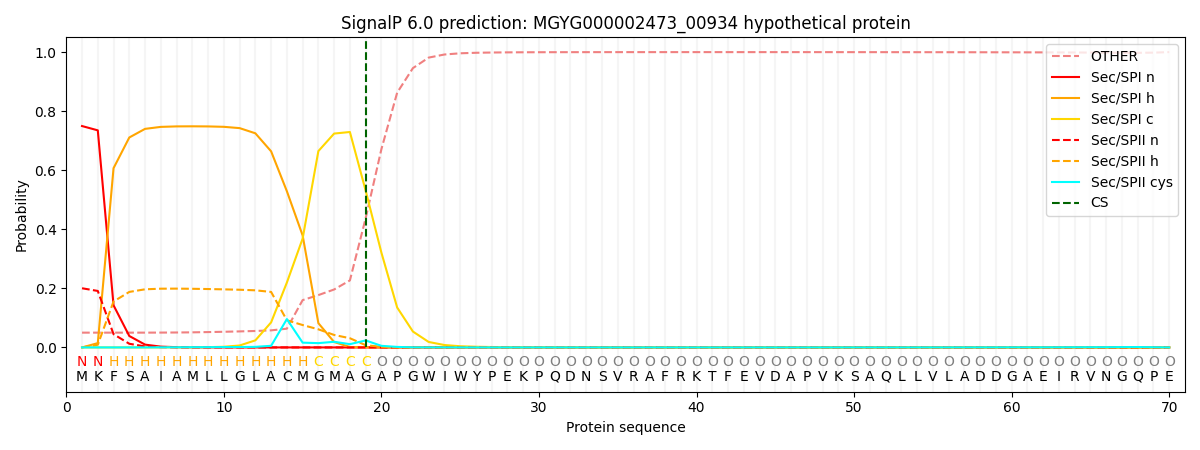
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 0.057395 | 0.733949 | 0.207378 | 0.000491 | 0.000349 | 0.000426 |