You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000002481_03276
You are here: Home > Sequence: MGYG000002481_03276
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Stenotrophomonas maltophilia_F | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Proteobacteria; Gammaproteobacteria; Xanthomonadales; Xanthomonadaceae; Stenotrophomonas; Stenotrophomonas maltophilia_F | |||||||||||
| CAZyme ID | MGYG000002481_03276 | |||||||||||
| CAZy Family | GT2 | |||||||||||
| CAZyme Description | Acetyl-coenzyme A synthetase | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 3669; End: 5612 Strand: + | |||||||||||
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| cd05967 | PrpE | 0.0 | 22 | 637 | 1 | 614 | Propionyl-CoA synthetase (PrpE). EC 6.2.1.17: propanoate:CoA ligase (AMP-forming) or propionate#CoA ligase (PrpE) catalyzes the first step of the 2-methylcitric acid cycle for propionate catabolism. It activates propionate to propionyl-CoA in a two-step reaction, which proceeds through a propionyl-AMP intermediate and requires ATP and Mg2+. In Salmonella enterica, the PrpE protein is required for growth of Salmonella enterica on propionate and can substitute for the acetyl-CoA synthetase (Acs) enzyme during growth on acetate. PrpE can also activate acetate, 3HP, and butyrate to their corresponding CoA-thioesters, although with less efficiency. |
| PLN02654 | PLN02654 | 0.0 | 4 | 641 | 14 | 661 | acetate-CoA ligase |
| PRK10524 | prpE | 0.0 | 20 | 635 | 2 | 618 | propionyl-CoA synthetase; Provisional |
| cd05968 | AACS_like | 0.0 | 15 | 634 | 2 | 610 | Uncharacterized acyl-CoA synthetase subfamily similar to Acetoacetyl-CoA synthetase. This uncharacterized acyl-CoA synthetase family (EC 6.2.1.16, or acetoacetate#CoA ligase or acetoacetate:CoA ligase (AMP-forming)) is highly homologous to acetoacetyl-CoA synthetase. However, the proteins in this family exist in only bacteria and archaea. AACS is a cytosolic ligase that specifically activates acetoacetate to its coenzyme A ester by a two-step reaction. Acetoacetate first reacts with ATP to form an acyl-adenylate intermediate, which then reacts with CoA to produce an acyl-CoA ester. This is the first step of the mevalonate pathway of isoprenoid biosynthesis via isopentenyl diphosphate. Isoprenoids are a large class of compounds found in all living organisms. |
| PRK00174 | PRK00174 | 0.0 | 4 | 645 | 1 | 634 | acetyl-CoA synthetase; Provisional |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| CAE6201330.1 | 9.54e-255 | 4 | 641 | 564 | 1210 |
| CAE6201338.1 | 1.32e-254 | 4 | 641 | 575 | 1221 |
| AFZ04852.1 | 6.30e-24 | 50 | 614 | 562 | 1074 |
| AFY93865.1 | 2.30e-23 | 83 | 640 | 340 | 867 |
| QND46664.1 | 2.23e-22 | 99 | 621 | 2086 | 2583 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 2P2F_A | 1.20e-311 | 5 | 645 | 7 | 646 | ChainA, Acetyl-coenzyme A synthetase [Salmonella enterica subsp. enterica serovar Typhimurium],2P2F_B Chain B, Acetyl-coenzyme A synthetase [Salmonella enterica subsp. enterica serovar Typhimurium] |
| 5JRH_A | 1.61e-311 | 5 | 645 | 7 | 646 | Crystalstructure of Salmonella enterica acetyl-CoA synthetase (Acs) in complex with cAMP and Coenzyme A [Salmonella enterica subsp. enterica serovar Typhimurium str. LT2],5JRH_B Crystal structure of Salmonella enterica acetyl-CoA synthetase (Acs) in complex with cAMP and Coenzyme A [Salmonella enterica subsp. enterica serovar Typhimurium str. LT2] |
| 2P2B_A | 4.88e-311 | 5 | 645 | 7 | 646 | ChainA, Acetyl-coenzyme A synthetase [Salmonella enterica subsp. enterica serovar Typhimurium],2P2B_B Chain B, Acetyl-coenzyme A synthetase [Salmonella enterica subsp. enterica serovar Typhimurium] |
| 2P2Q_A | 6.93e-311 | 5 | 645 | 7 | 646 | ChainA, Acetyl-coenzyme A synthetase [Salmonella enterica subsp. enterica serovar Typhimurium],2P2Q_B Chain B, Acetyl-coenzyme A synthetase [Salmonella enterica subsp. enterica serovar Typhimurium] |
| 2P2J_A | 9.83e-311 | 5 | 645 | 7 | 646 | ChainA, Acetyl-coenzyme A synthetase [Salmonella enterica subsp. enterica serovar Typhimurium],2P2J_B Chain B, Acetyl-coenzyme A synthetase [Salmonella enterica subsp. enterica serovar Typhimurium] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| Q2YLH5 | 0.0 | 4 | 645 | 5 | 648 | Acetyl-coenzyme A synthetase OS=Brucella abortus (strain 2308) OX=359391 GN=acsA PE=3 SV=1 |
| Q87C00 | 0.0 | 1 | 644 | 1 | 644 | Acetyl-coenzyme A synthetase OS=Xylella fastidiosa (strain Temecula1 / ATCC 700964) OX=183190 GN=acsA PE=3 SV=1 |
| Q6MNF1 | 0.0 | 3 | 647 | 4 | 645 | Acetyl-coenzyme A synthetase OS=Bdellovibrio bacteriovorus (strain ATCC 15356 / DSM 50701 / NCIMB 9529 / HD100) OX=264462 GN=acsA PE=3 SV=1 |
| Q4UP35 | 0.0 | 1 | 647 | 1 | 647 | Acetyl-coenzyme A synthetase OS=Xanthomonas campestris pv. campestris (strain 8004) OX=314565 GN=acsA PE=3 SV=1 |
| Q89WV5 | 0.0 | 4 | 645 | 5 | 643 | Acetyl-coenzyme A synthetase OS=Bradyrhizobium diazoefficiens (strain JCM 10833 / BCRC 13528 / IAM 13628 / NBRC 14792 / USDA 110) OX=224911 GN=acsA PE=1 SV=2 |
SignalP and Lipop Annotations help
This protein is predicted as OTHER
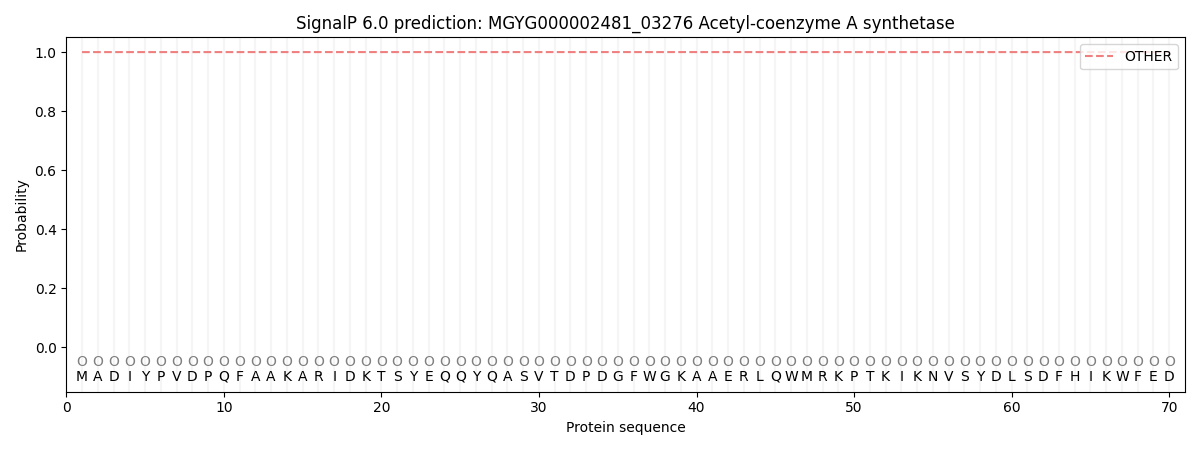
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 1.000058 | 0.000000 | 0.000000 | 0.000000 | 0.000000 | 0.000000 |
