You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000002492_02420
You are here: Home > Sequence: MGYG000002492_02420
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Agathobacter rectalis | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Firmicutes_A; Clostridia; Lachnospirales; Lachnospiraceae; Agathobacter; Agathobacter rectalis | |||||||||||
| CAZyme ID | MGYG000002492_02420 | |||||||||||
| CAZy Family | GT1 | |||||||||||
| CAZyme Description | hypothetical protein | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 2537021; End: 2537479 Strand: - | |||||||||||
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| pfam08660 | Alg14 | 1.11e-22 | 6 | 149 | 1 | 169 | Oligosaccharide biosynthesis protein Alg14 like. Alg14 is involved dolichol-linked oligosaccharide biosynthesis and anchors the catalytic subunit Alg13 to the ER membrane. |
| COG0707 | MurG | 5.64e-12 | 9 | 145 | 7 | 154 | UDP-N-acetylglucosamine:LPS N-acetylglucosamine transferase [Cell wall/membrane/envelope biogenesis]. |
| cd03785 | GT28_MurG | 1.31e-05 | 59 | 145 | 69 | 152 | undecaprenyldiphospho-muramoylpentapeptide beta-N-acetylglucosaminyltransferase. MurG (EC 2.4.1.227) is an N-acetylglucosaminyltransferase, the last enzyme involved in the intracellular phase of peptidoglycan biosynthesis. It transfers N-acetyl-D-glucosamine (GlcNAc) from UDP-GlcNAc to the C4 hydroxyl of a lipid-linked N-acetylmuramoyl pentapeptide (NAM). The resulting disaccharide is then transported across the cell membrane, where it is polymerized into NAG-NAM cell-wall repeat structure. MurG belongs to the GT-B structural superfamily of glycoslytransferases, which have characteristic N- and C-terminal domains, each containing a typical Rossmann fold. The two domains have high structural homology despite minimal sequence homology. The large cleft that separates the two domains includes the catalytic center and permits a high degree of flexibility. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| ACR76392.1 | 3.32e-112 | 1 | 152 | 1 | 152 |
| ABJ66265.1 | 5.79e-103 | 2 | 152 | 8 | 158 |
| AIR11263.1 | 8.08e-95 | 1 | 152 | 1 | 152 |
| ASM70738.1 | 2.97e-94 | 4 | 152 | 1 | 149 |
| QGF40721.1 | 4.22e-94 | 4 | 152 | 1 | 149 |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| Q96F25 | 3.33e-12 | 2 | 151 | 35 | 215 | UDP-N-acetylglucosamine transferase subunit ALG14 homolog OS=Homo sapiens OX=9606 GN=ALG14 PE=1 SV=1 |
| Q9D081 | 3.25e-09 | 2 | 151 | 35 | 216 | UDP-N-acetylglucosamine transferase subunit ALG14 homolog OS=Mus musculus OX=10090 GN=Alg14 PE=1 SV=1 |
| Q5A5N6 | 4.59e-09 | 60 | 151 | 121 | 218 | UDP-N-acetylglucosamine transferase subunit ALG14 OS=Candida albicans (strain SC5314 / ATCC MYA-2876) OX=237561 GN=ALG14 PE=3 SV=1 |
| Q6CJG3 | 7.38e-09 | 61 | 151 | 137 | 235 | UDP-N-acetylglucosamine transferase subunit ALG14 OS=Kluyveromyces lactis (strain ATCC 8585 / CBS 2359 / DSM 70799 / NBRC 1267 / NRRL Y-1140 / WM37) OX=284590 GN=ALG14 PE=3 SV=1 |
| O14199 | 8.00e-09 | 80 | 151 | 133 | 209 | UDP-N-acetylglucosamine transferase subunit alg14 OS=Schizosaccharomyces pombe (strain 972 / ATCC 24843) OX=284812 GN=alg14 PE=3 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as OTHER
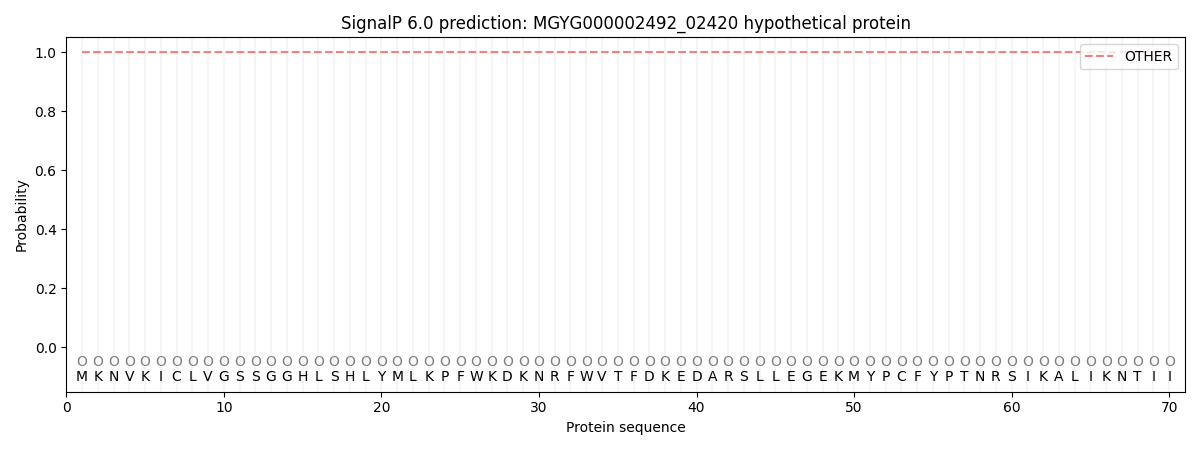
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 1.000077 | 0.000000 | 0.000000 | 0.000000 | 0.000000 | 0.000000 |
