You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000002514_02258
You are here: Home > Sequence: MGYG000002514_02258
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
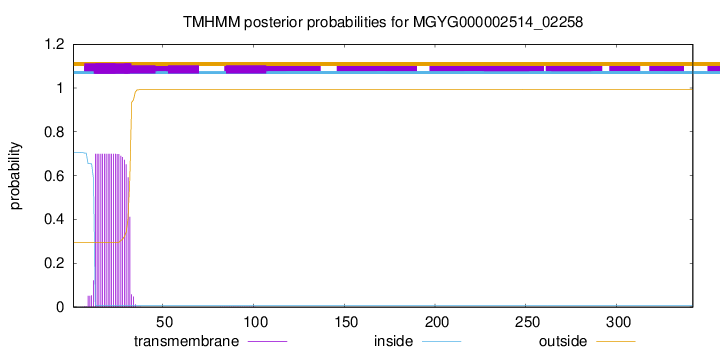
TMHMM annotations
Basic Information help
| Species | Proteus mirabilis | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Proteobacteria; Gammaproteobacteria; Enterobacterales; Enterobacteriaceae; Proteus; Proteus mirabilis | |||||||||||
| CAZyme ID | MGYG000002514_02258 | |||||||||||
| CAZy Family | GH8 | |||||||||||
| CAZyme Description | Minor endoglucanase Y | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 2410233; End: 2411261 Strand: + | |||||||||||
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| pfam01270 | Glyco_hydro_8 | 7.60e-94 | 37 | 334 | 7 | 314 | Glycosyl hydrolases family 8. |
| COG3405 | BcsZ | 2.59e-62 | 16 | 341 | 8 | 343 | Endo-1,4-beta-D-glucanase Y [Carbohydrate transport and metabolism]. |
| PRK11097 | PRK11097 | 9.37e-44 | 17 | 277 | 7 | 279 | cellulase. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| AWR59912.1 | 7.75e-260 | 1 | 342 | 1 | 342 |
| AZH00392.1 | 2.90e-258 | 1 | 342 | 1 | 342 |
| QIG03487.1 | 5.24e-258 | 1 | 342 | 1 | 342 |
| QOV82867.1 | 5.24e-258 | 1 | 342 | 1 | 342 |
| QWV42057.1 | 5.24e-258 | 1 | 342 | 1 | 342 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 5GY3_A | 2.37e-102 | 35 | 341 | 1 | 308 | ChainA, Glucanase [Klebsiella pneumoniae] |
| 5CZL_A | 4.19e-102 | 35 | 341 | 29 | 336 | ChainA, Glucanase [Raoultella ornithinolytica] |
| 6VC5_A | 2.39e-75 | 38 | 340 | 8 | 314 | 1.6Angstrom Resolution Crystal Structure of endoglucanase from Komagataeibacter sucrofermentans [Komagataeibacter sucrofermentans] |
| 1WZZ_A | 3.54e-72 | 38 | 339 | 23 | 328 | Structureof endo-beta-1,4-glucanase CMCax from Acetobacter xylinum [Komagataeibacter xylinus] |
| 4Q2B_A | 3.27e-31 | 38 | 259 | 7 | 230 | Thecrystal structure of an endo-1,4-D-glucanase from Pseudomonas putida KT2440 [Pseudomonas putida KT2440],4Q2B_B The crystal structure of an endo-1,4-D-glucanase from Pseudomonas putida KT2440 [Pseudomonas putida KT2440],4Q2B_C The crystal structure of an endo-1,4-D-glucanase from Pseudomonas putida KT2440 [Pseudomonas putida KT2440],4Q2B_D The crystal structure of an endo-1,4-D-glucanase from Pseudomonas putida KT2440 [Pseudomonas putida KT2440],4Q2B_E The crystal structure of an endo-1,4-D-glucanase from Pseudomonas putida KT2440 [Pseudomonas putida KT2440],4Q2B_F The crystal structure of an endo-1,4-D-glucanase from Pseudomonas putida KT2440 [Pseudomonas putida KT2440] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| P27032 | 3.15e-110 | 23 | 339 | 11 | 330 | Minor endoglucanase Y OS=Dickeya dadantii (strain 3937) OX=198628 GN=celY PE=1 SV=1 |
| P18336 | 9.48e-98 | 35 | 321 | 24 | 311 | Endoglucanase OS=Cellulomonas uda OX=1714 PE=1 SV=1 |
| P37696 | 3.89e-73 | 38 | 339 | 31 | 336 | Probable endoglucanase OS=Komagataeibacter hansenii OX=436 GN=cmcAX PE=1 SV=1 |
| Q8X5L9 | 4.66e-29 | 30 | 264 | 20 | 256 | Endoglucanase OS=Escherichia coli O157:H7 OX=83334 GN=bcsZ PE=3 SV=1 |
| P37651 | 6.45e-29 | 30 | 264 | 20 | 256 | Endoglucanase OS=Escherichia coli (strain K12) OX=83333 GN=bcsZ PE=1 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as SP
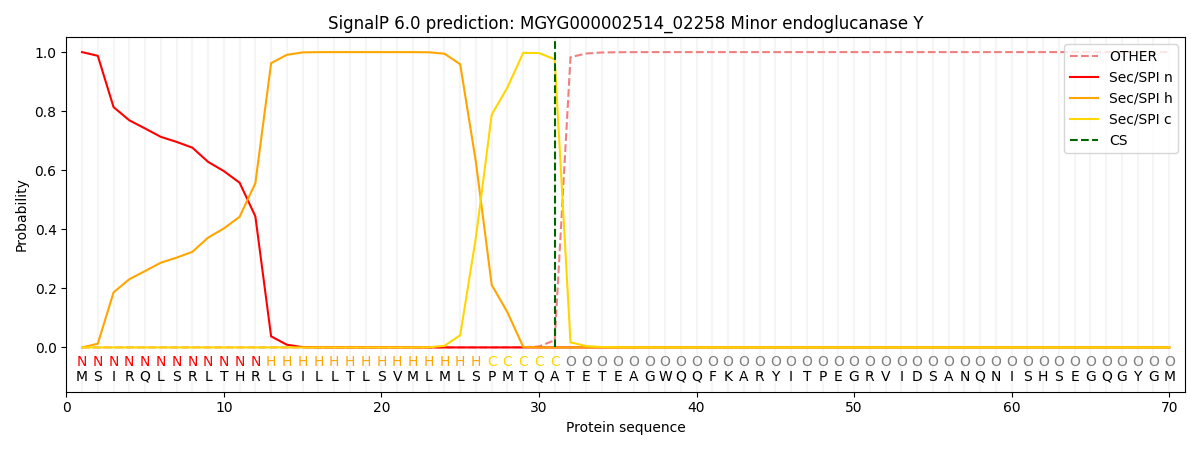
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 0.000261 | 0.999035 | 0.000190 | 0.000184 | 0.000157 | 0.000143 |