You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000002517_01134
You are here: Home > Sequence: MGYG000002517_01134
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Roseburia hominis | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Firmicutes_A; Clostridia; Lachnospirales; Lachnospiraceae; Roseburia; Roseburia hominis | |||||||||||
| CAZyme ID | MGYG000002517_01134 | |||||||||||
| CAZy Family | CE17 | |||||||||||
| CAZyme Description | hypothetical protein | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 1247868; End: 1249601 Strand: + | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| CE17 | 215 | 384 | 6.2e-43 | 0.9878787878787879 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| cd00229 | SGNH_hydrolase | 1.02e-17 | 213 | 391 | 1 | 184 | SGNH_hydrolase, or GDSL_hydrolase, is a diverse family of lipases and esterases. The tertiary fold of the enzyme is substantially different from that of the alpha/beta hydrolase family and unique among all known hydrolases; its active site closely resembles the typical Ser-His-Asp(Glu) triad from other serine hydrolases, but may lack the carboxlic acid. |
| pfam13472 | Lipase_GDSL_2 | 2.73e-15 | 215 | 386 | 1 | 176 | GDSL-like Lipase/Acylhydrolase family. This family of presumed lipases and related enzymes are similar to pfam00657. |
| cd01834 | SGNH_hydrolase_like_2 | 9.02e-11 | 213 | 388 | 4 | 185 | SGNH_hydrolase subfamily. SGNH hydrolases are a diverse family of lipases and esterases. The tertiary fold of the enzyme is substantially different from that of the alpha/beta hydrolase family and unique among all known hydrolases; its active site closely resembles the Ser-His-Asp(Glu) triad found in other serine hydrolases. |
| COG2755 | TesA | 8.89e-07 | 212 | 389 | 10 | 202 | Lysophospholipase L1 or related esterase [Amino acid transport and metabolism]. |
| cd01827 | sialate_O-acetylesterase_like1 | 7.15e-05 | 211 | 388 | 1 | 180 | sialate O-acetylesterase_like family of the SGNH hydrolases, a diverse family of lipases and esterases. The tertiary fold of the enzyme is substantially different from that of the alpha/beta hydrolase family and unique among all known hydrolases; its active site closely resembles the Ser-His-Asp(Glu) triad found in other serine hydrolases. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| QSF46877.1 | 1.66e-195 | 2 | 575 | 14 | 577 |
| QYK61360.1 | 1.97e-194 | 1 | 576 | 14 | 579 |
| AEN97394.1 | 2.83e-40 | 185 | 396 | 7 | 211 |
| EEV02614.1 | 2.54e-38 | 180 | 402 | 2 | 217 |
| CBL10432.1 | 3.50e-38 | 180 | 402 | 2 | 217 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 6HH9_A | 1.05e-39 | 180 | 402 | 2 | 217 | Crystalstructure of a two-domain esterase (CEX) active on acetylated mannans co-crystallized with mannopentaose [Roseburia intestinalis L1-82],6HH9_B Crystal structure of a two-domain esterase (CEX) active on acetylated mannans co-crystallized with mannopentaose [Roseburia intestinalis L1-82],6HH9_C Crystal structure of a two-domain esterase (CEX) active on acetylated mannans co-crystallized with mannopentaose [Roseburia intestinalis L1-82],6HH9_D Crystal structure of a two-domain esterase (CEX) active on acetylated mannans co-crystallized with mannopentaose [Roseburia intestinalis L1-82] |
| 6HFZ_A | 2.00e-39 | 180 | 402 | 2 | 217 | Crystalstructure of a two-domain esterase (CEX) active on acetylated mannans [Roseburia intestinalis L1-82],6HFZ_B Crystal structure of a two-domain esterase (CEX) active on acetylated mannans [Roseburia intestinalis L1-82],6HFZ_C Crystal structure of a two-domain esterase (CEX) active on acetylated mannans [Roseburia intestinalis L1-82],6HFZ_D Crystal structure of a two-domain esterase (CEX) active on acetylated mannans [Roseburia intestinalis L1-82] |
Swiss-Prot Hits help
SignalP and Lipop Annotations help
This protein is predicted as OTHER
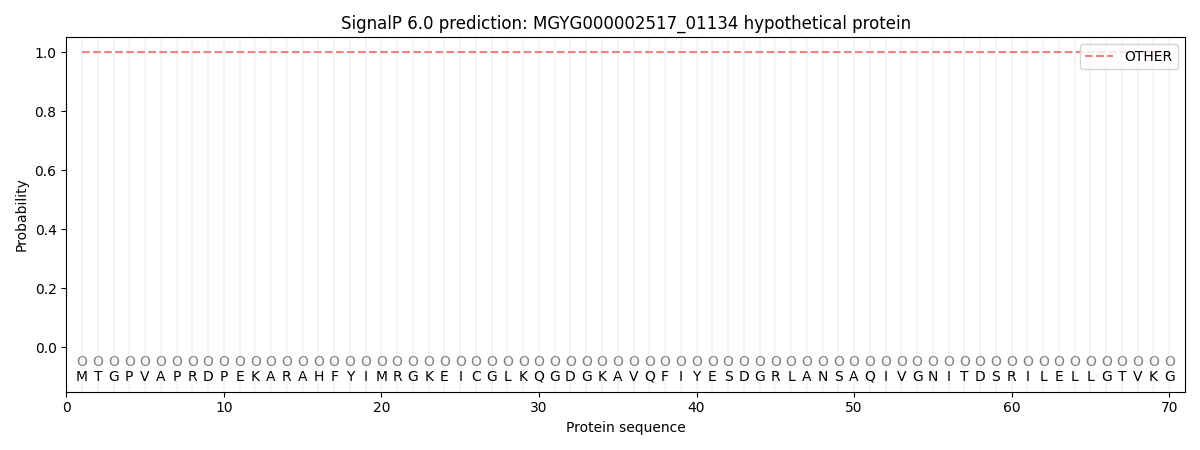
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 1.000077 | 0.000000 | 0.000000 | 0.000000 | 0.000000 | 0.000000 |
