You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000002521_05403
You are here: Home > Sequence: MGYG000002521_05403
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Phytobacter ursingii | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Proteobacteria; Gammaproteobacteria; Enterobacterales; Enterobacteriaceae; Phytobacter; Phytobacter ursingii | |||||||||||
| CAZyme ID | MGYG000002521_05403 | |||||||||||
| CAZy Family | GH73 | |||||||||||
| CAZyme Description | hypothetical protein | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 4832606; End: 4833433 Strand: + | |||||||||||
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| PRK10356 | PRK10356 | 1.87e-175 | 1 | 275 | 1 | 274 | protein bax. |
| COG2992 | Bax | 2.55e-125 | 13 | 275 | 2 | 262 | Uncharacterized FlgJ-related protein [General function prediction only]. |
| smart00047 | LYZ2 | 1.75e-27 | 128 | 257 | 1 | 147 | Lysozyme subfamily 2. Eubacterial enzymes distantly related to eukaryotic lysozymes. |
| pfam01832 | Glucosaminidase | 6.66e-14 | 142 | 210 | 6 | 77 | Mannosyl-glycoprotein endo-beta-N-acetylglucosaminidase. This family includes Mannosyl-glycoprotein endo-beta-N-acetylglucosaminidase EC:3.2.1.96. As well as the flageller protein J that has been shown to hydrolyze peptidoglycan. |
| NF038016 | sporang_Gsm | 1.41e-06 | 157 | 182 | 185 | 212 | sporangiospore maturation cell wall hydrolase GsmA. The peptidoglycan-hydrolyzing enzyme GsmA occurs in some sporangia-forming members of the Actinobacteria, such as Actinoplanes missouriensis, and is required for proper separation of spores. GsmA proteins have one or two SH3 domains N-terminal to the hydrolase domain. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| AKL14577.1 | 1.51e-196 | 1 | 275 | 1 | 275 |
| AUV03561.1 | 8.75e-196 | 1 | 275 | 1 | 275 |
| QJF15576.1 | 9.79e-193 | 1 | 275 | 1 | 275 |
| QOV68692.1 | 9.79e-193 | 1 | 275 | 1 | 275 |
| QIH66203.1 | 9.79e-193 | 1 | 275 | 1 | 275 |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| P27297 | 2.73e-154 | 1 | 275 | 1 | 274 | Protein bax OS=Escherichia coli (strain K12) OX=83333 GN=bax PE=4 SV=3 |
SignalP and Lipop Annotations help
This protein is predicted as SP
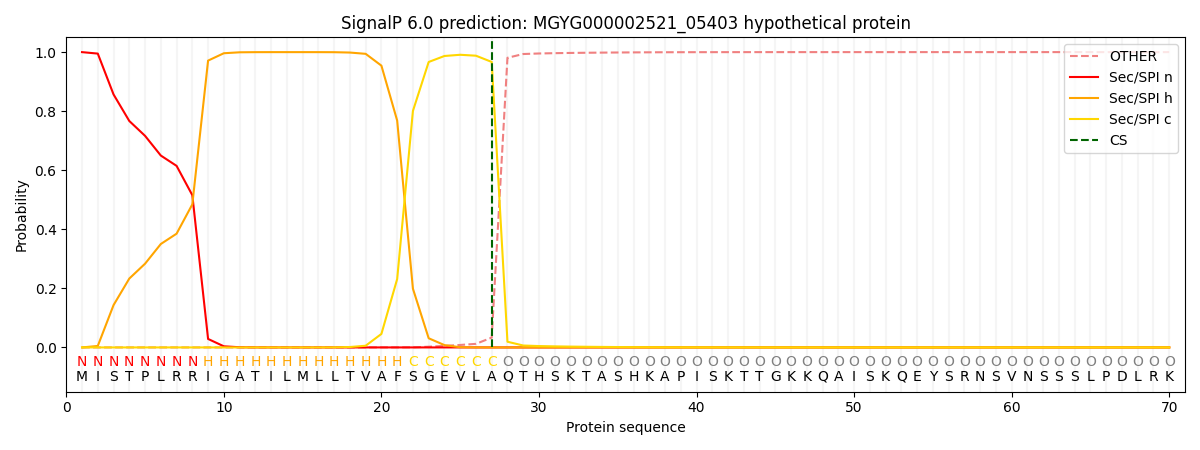
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 0.000710 | 0.998391 | 0.000333 | 0.000182 | 0.000176 | 0.000166 |
