You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000002531_02288
You are here: Home > Sequence: MGYG000002531_02288
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Pseudobutyrivibrio fibrisolvens_A | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Firmicutes_A; Clostridia; Lachnospirales; Lachnospiraceae; Pseudobutyrivibrio; Pseudobutyrivibrio fibrisolvens_A | |||||||||||
| CAZyme ID | MGYG000002531_02288 | |||||||||||
| CAZy Family | GH10 | |||||||||||
| CAZyme Description | hypothetical protein | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 2171696; End: 2173768 Strand: - | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH10 | 42 | 381 | 1.5e-80 | 0.9801980198019802 |
| CBM2 | 604 | 681 | 4.4e-24 | 0.7722772277227723 |
| CBM13 | 427 | 571 | 4.5e-23 | 0.7127659574468085 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| smart00633 | Glyco_10 | 6.50e-71 | 116 | 378 | 12 | 263 | Glycosyl hydrolase family 10. |
| pfam00331 | Glyco_hydro_10 | 1.75e-64 | 48 | 378 | 11 | 308 | Glycosyl hydrolase family 10. |
| COG3693 | XynA | 3.75e-51 | 11 | 375 | 3 | 334 | Endo-1,4-beta-xylanase, GH35 family [Carbohydrate transport and metabolism]. |
| smart00637 | CBD_II | 2.74e-20 | 606 | 685 | 2 | 82 | CBD_II domain. |
| pfam00553 | CBM_2 | 5.47e-20 | 603 | 681 | 6 | 85 | Cellulose binding domain. Two tryptophan residues are involved in cellulose binding. Cellulose binding domain found in bacteria. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| CBK75021.1 | 0.0 | 1 | 690 | 1 | 690 |
| QFJ54853.1 | 0.0 | 1 | 690 | 1 | 694 |
| ADL33480.1 | 0.0 | 35 | 683 | 707 | 1386 |
| ADL33750.1 | 2.79e-294 | 1 | 684 | 3 | 683 |
| AQS12857.1 | 9.03e-200 | 30 | 569 | 30 | 561 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 2W5F_A | 1.57e-48 | 30 | 393 | 176 | 539 | ChainA, ENDO-1,4-BETA-XYLANASE Y [Acetivibrio thermocellus],2W5F_B Chain B, ENDO-1,4-BETA-XYLANASE Y [Acetivibrio thermocellus] |
| 2WYS_A | 8.41e-46 | 30 | 393 | 176 | 539 | ChainA, ENDO-1,4-BETA-XYLANASE Y [Acetivibrio thermocellus],2WYS_B Chain B, ENDO-1,4-BETA-XYLANASE Y [Acetivibrio thermocellus],2WZE_A Chain A, ENDO-1,4-BETA-XYLANASE Y [Acetivibrio thermocellus],2WZE_B Chain B, ENDO-1,4-BETA-XYLANASE Y [Acetivibrio thermocellus] |
| 5OFJ_A | 3.08e-38 | 37 | 381 | 10 | 338 | Crystalstructure of N-terminal domain of bifunctional CbXyn10C [Caldicellulosiruptor bescii DSM 6725] |
| 6D5C_A | 5.37e-38 | 69 | 381 | 53 | 350 | Structureof Caldicellulosiruptor danielii GH10 module of glycoside hydrolase WP_045175321 [Caldicellulosiruptor danielii],6D5C_B Structure of Caldicellulosiruptor danielii GH10 module of glycoside hydrolase WP_045175321 [Caldicellulosiruptor danielii],6D5C_C Structure of Caldicellulosiruptor danielii GH10 module of glycoside hydrolase WP_045175321 [Caldicellulosiruptor danielii] |
| 6FHE_A | 5.75e-38 | 69 | 379 | 44 | 339 | Highlyactive enzymes by automated modular backbone assembly and sequence design [synthetic construct] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| P23551 | 2.18e-185 | 1 | 391 | 1 | 392 | Endo-1,4-beta-xylanase A OS=Butyrivibrio fibrisolvens OX=831 GN=xynA PE=3 SV=1 |
| P29126 | 2.12e-53 | 50 | 379 | 641 | 950 | Bifunctional endo-1,4-beta-xylanase XylA OS=Ruminococcus flavefaciens OX=1265 GN=xynA PE=3 SV=1 |
| P51584 | 8.43e-47 | 30 | 393 | 187 | 550 | Endo-1,4-beta-xylanase Y OS=Acetivibrio thermocellus OX=1515 GN=xynY PE=1 SV=1 |
| P26223 | 3.62e-41 | 39 | 381 | 3 | 337 | Endo-1,4-beta-xylanase B OS=Butyrivibrio fibrisolvens OX=831 GN=xynB PE=3 SV=1 |
| P40944 | 6.20e-36 | 37 | 379 | 355 | 676 | Endo-1,4-beta-xylanase A OS=Caldicellulosiruptor sp. (strain Rt8B.4) OX=28238 GN=xynA PE=3 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as SP
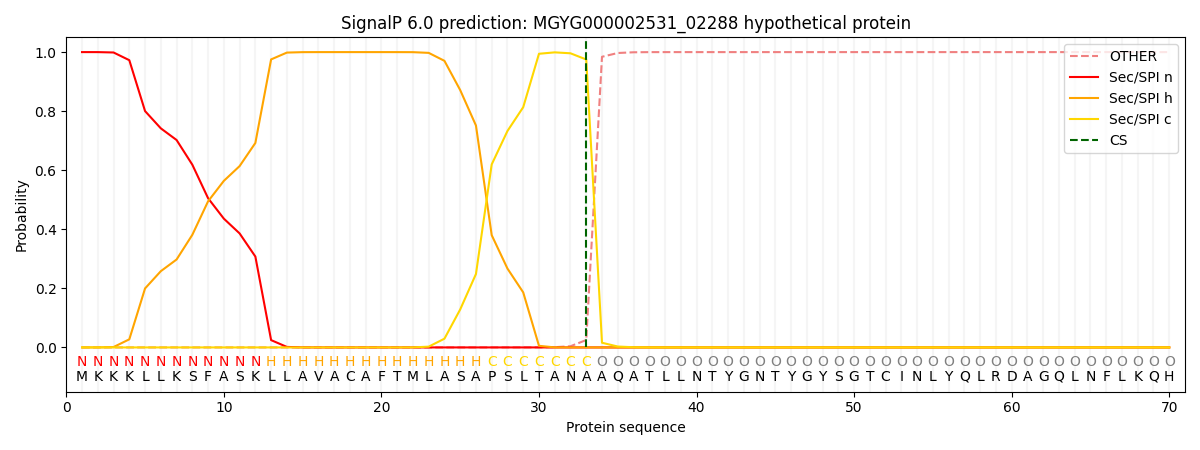
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 0.000249 | 0.999051 | 0.000180 | 0.000186 | 0.000163 | 0.000145 |
