You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000002558_00800
You are here: Home > Sequence: MGYG000002558_00800
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Prevotella sp900540415 | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Bacteroidota; Bacteroidia; Bacteroidales; Bacteroidaceae; Prevotella; Prevotella sp900540415 | |||||||||||
| CAZyme ID | MGYG000002558_00800 | |||||||||||
| CAZy Family | CE8 | |||||||||||
| CAZyme Description | Pectinesterase A | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 28036; End: 29022 Strand: + | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| CE8 | 30 | 315 | 1.6e-103 | 0.9861111111111112 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| pfam01095 | Pectinesterase | 2.33e-83 | 30 | 310 | 2 | 290 | Pectinesterase. |
| PLN02773 | PLN02773 | 2.25e-70 | 32 | 307 | 9 | 288 | pectinesterase |
| PLN02197 | PLN02197 | 1.36e-66 | 29 | 311 | 276 | 567 | pectinesterase |
| COG4677 | PemB | 3.69e-64 | 30 | 315 | 84 | 402 | Pectin methylesterase and related acyl-CoA thioesterases [Carbohydrate transport and metabolism, Lipid transport and metabolism]. |
| PLN02682 | PLN02682 | 1.55e-63 | 22 | 306 | 63 | 352 | pectinesterase family protein |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| QVJ81978.1 | 1.18e-175 | 3 | 323 | 2 | 322 |
| ADE82952.1 | 1.18e-175 | 3 | 323 | 2 | 322 |
| BCS84477.1 | 9.75e-164 | 3 | 327 | 2 | 327 |
| QQY38333.1 | 5.94e-160 | 25 | 323 | 21 | 314 |
| ALK82867.1 | 5.94e-160 | 25 | 323 | 21 | 314 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 1GQ8_A | 2.16e-48 | 30 | 310 | 9 | 297 | Pectinmethylesterase from Carrot [Daucus carota] |
| 5C1E_A | 3.51e-44 | 30 | 294 | 11 | 271 | CrystalStructure of the Pectin Methylesterase from Aspergillus niger in Penultimately Deglycosylated Form (N-acetylglucosamine Stub at Asn84) [Aspergillus niger ATCC 1015] |
| 1XG2_A | 5.48e-44 | 31 | 297 | 6 | 277 | ChainA, Pectinesterase 1 [Solanum lycopersicum] |
| 5C1C_A | 1.91e-43 | 30 | 294 | 11 | 271 | CrystalStructure of the Pectin Methylesterase from Aspergillus niger in Deglycosylated Form [Aspergillus niger ATCC 1015] |
| 3UW0_A | 1.15e-34 | 59 | 315 | 62 | 358 | Pectinmethylesterase from Yersinia enterocolitica [Yersinia enterocolitica subsp. enterocolitica 8081] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| P41510 | 2.06e-60 | 29 | 325 | 272 | 576 | Probable pectinesterase/pectinesterase inhibitor OS=Brassica napus OX=3708 GN=BP19 PE=2 SV=1 |
| Q42608 | 3.46e-58 | 29 | 325 | 259 | 563 | Pectinesterase/pectinesterase inhibitor (Fragment) OS=Brassica campestris OX=3711 PE=2 SV=1 |
| Q5MFV6 | 6.55e-58 | 29 | 307 | 276 | 561 | Probable pectinesterase/pectinesterase inhibitor VGDH2 OS=Arabidopsis thaliana OX=3702 GN=VGDH2 PE=2 SV=2 |
| O80722 | 2.65e-55 | 31 | 310 | 278 | 566 | Pectinesterase 4 OS=Arabidopsis thaliana OX=3702 GN=PME4 PE=2 SV=1 |
| Q9LVQ0 | 1.19e-54 | 27 | 306 | 4 | 287 | Pectinesterase 31 OS=Arabidopsis thaliana OX=3702 GN=PME31 PE=1 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as SP
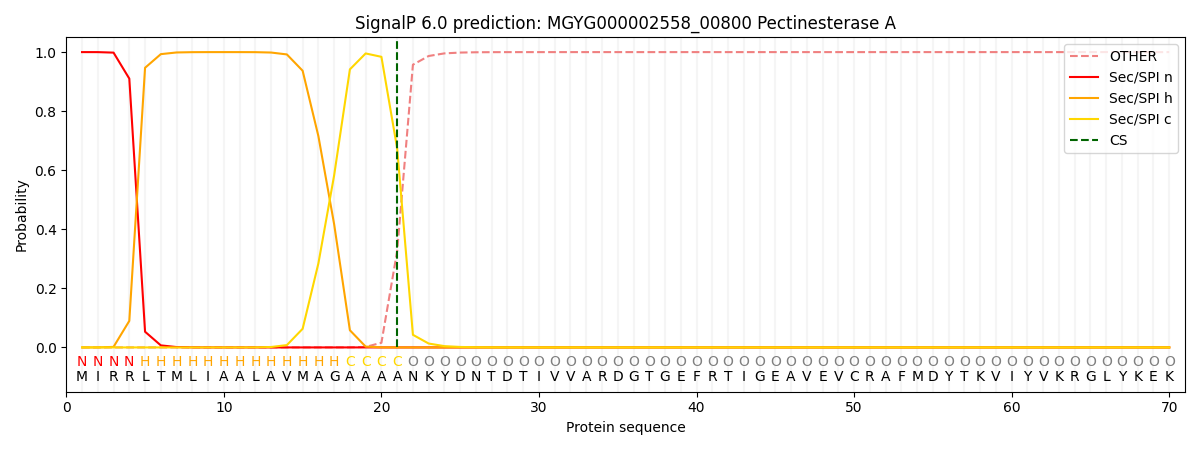
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 0.000495 | 0.998785 | 0.000189 | 0.000184 | 0.000167 | 0.000162 |
