You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000002561_02521
You are here: Home > Sequence: MGYG000002561_02521
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Bacteroides sp902388495 | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Bacteroidota; Bacteroidia; Bacteroidales; Bacteroidaceae; Bacteroides; Bacteroides sp902388495 | |||||||||||
| CAZyme ID | MGYG000002561_02521 | |||||||||||
| CAZy Family | GH55 | |||||||||||
| CAZyme Description | hypothetical protein | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 11680; End: 14694 Strand: - | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH55 | 38 | 673 | 5.3e-89 | 0.8581081081081081 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| COG3386 | YvrE | 5.48e-13 | 703 | 994 | 2 | 274 | Sugar lactone lactonase YvrE [Carbohydrate transport and metabolism]. |
| pfam12708 | Pectate_lyase_3 | 6.42e-12 | 45 | 217 | 7 | 171 | Pectate lyase superfamily protein. This family of proteins possesses a beta helical structure like Pectate lyase. This family is most closely related to glycosyl hydrolase family 28. |
| pfam12708 | Pectate_lyase_3 | 2.65e-11 | 378 | 431 | 1 | 60 | Pectate lyase superfamily protein. This family of proteins possesses a beta helical structure like Pectate lyase. This family is most closely related to glycosyl hydrolase family 28. |
| pfam08450 | SGL | 2.83e-11 | 905 | 993 | 145 | 243 | SMP-30/Gluconolaconase/LRE-like region. This family describes a region that is found in proteins expressed by a variety of eukaryotic and prokaryotic species. These proteins include various enzymes, such as senescence marker protein 30 (SMP-30), gluconolactonase and luciferin-regenerating enzyme (LRE). SMP-30 is known to hydrolyze diisopropyl phosphorofluoridate in the liver, and has been noted as having sequence similarity, in the region described in this family, with PON1 and LRE. |
| COG5434 | Pgu1 | 3.17e-08 | 313 | 489 | 20 | 222 | Polygalacturonase [Carbohydrate transport and metabolism]. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| QUU00135.1 | 0.0 | 1 | 1004 | 1 | 1005 |
| QMI79027.1 | 0.0 | 1 | 1004 | 1 | 1004 |
| QPH57852.1 | 0.0 | 1 | 1004 | 1 | 1004 |
| QBJ17509.1 | 0.0 | 1 | 1004 | 1 | 1004 |
| QUT68914.1 | 0.0 | 1 | 1004 | 1 | 1004 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 5M5Z_A | 1.25e-06 | 365 | 431 | 374 | 439 | ChainA, Beta-1,3-glucanase [Thermochaetoides thermophila],5M60_A Chain A, Beta-1,3-glucanase [Thermochaetoides thermophila] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| D4B0V1 | 1.85e-06 | 378 | 431 | 511 | 567 | Probable glucan endo-1,3-beta-glucosidase ARB_02077 OS=Arthroderma benhamiae (strain ATCC MYA-4681 / CBS 112371) OX=663331 GN=ARB_02077 PE=1 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as SP
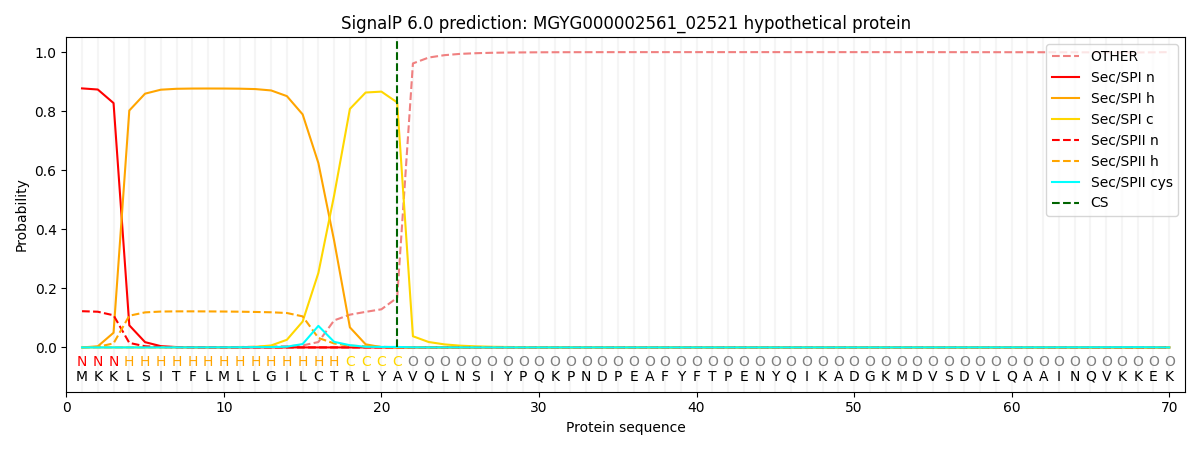
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 0.001634 | 0.870266 | 0.127245 | 0.000283 | 0.000278 | 0.000260 |
