You are browsing environment: HUMAN GUT
MGYG000002568_01613
Basic Information
help
Species
Phocaeicola sp900542985
Lineage
Bacteria; Bacteroidota; Bacteroidia; Bacteroidales; Bacteroidaceae; Phocaeicola; Phocaeicola sp900542985
CAZyme ID
MGYG000002568_01613
CAZy Family
GH163
CAZyme Description
hypothetical protein
CAZyme Property
Genome Property
Genome Assembly ID
Genome Size
Genome Type
Country
Continent
MGYG000002568
2891879
MAG
China
Asia
Gene Location
Start: 24476;
End: 26668
Strand: -
No EC number prediction in MGYG000002568_01613.
Family
Start
End
Evalue
family coverage
GH163
195
447
5.2e-107
0.9960159362549801
Cdd ID
Domain
E-Value
qStart
qEnd
sStart
sEnd
Domain Description
pfam16126
DUF4838
2.74e-131
188
447
1
263
Domain of unknown function (DUF4838). This family consists of several uncharacterized proteins found in various Bacteroides and Chloroflexus species. The function of this family is unknown.
more
pfam00754
F5_F8_type_C
6.35e-07
595
705
13
114
F5/8 type C domain. This domain is also known as the discoidin (DS) domain family.
more
pfam02838
Glyco_hydro_20b
6.67e-06
32
119
21
105
Glycosyl hydrolase family 20, domain 2. This domain has a zincin-like fold.
more
Hit ID
E-Value
Query Start
Query End
Hit Start
Hit End
Description
6Q63_A
5.62e-11
587
730
629
774
BT0459[Bacteroides thetaiotaomicron],6Q63_B BT0459 [Bacteroides thetaiotaomicron],6Q63_C BT0459 [Bacteroides thetaiotaomicron]
more
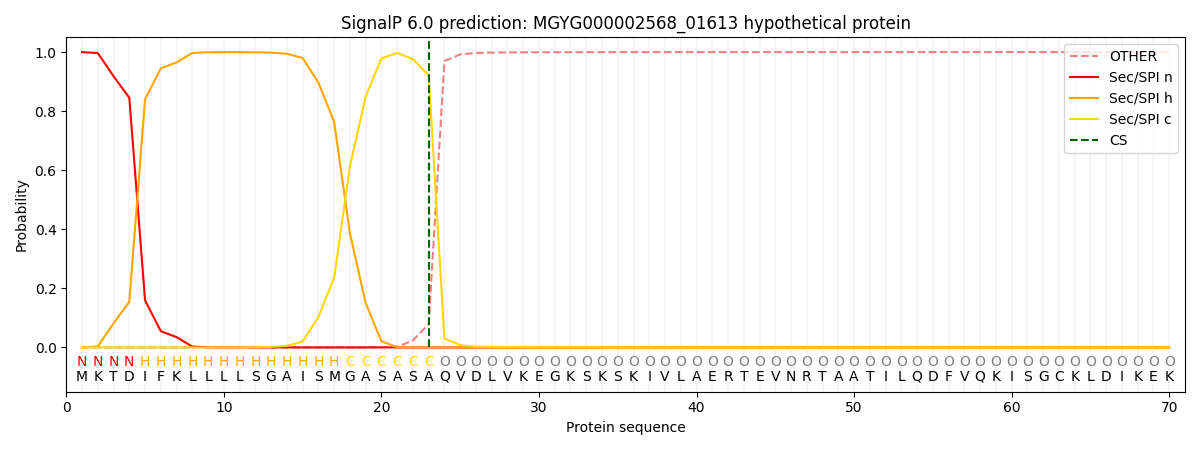
This protein is predicted as SP
Other
SP_Sec_SPI
LIPO_Sec_SPII
TAT_Tat_SPI
TATLIP_Sec_SPII
PILIN_Sec_SPIII
0.000482
0.998784
0.000184
0.000190
0.000167
0.000157
There is no transmembrane helices in MGYG000002568_01613.