You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000002579_00333
You are here: Home > Sequence: MGYG000002579_00333
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Ligilactobacillus sp900765635 | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Firmicutes; Bacilli; Lactobacillales; Lactobacillaceae; Ligilactobacillus; Ligilactobacillus sp900765635 | |||||||||||
| CAZyme ID | MGYG000002579_00333 | |||||||||||
| CAZy Family | GH29 | |||||||||||
| CAZyme Description | hypothetical protein | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 68050; End: 69060 Strand: + | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH29 | 3 | 334 | 5.5e-84 | 0.8526011560693642 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| pfam01120 | Alpha_L_fucos | 2.24e-78 | 1 | 328 | 18 | 323 | Alpha-L-fucosidase. |
| smart00812 | Alpha_L_fucos | 4.79e-73 | 1 | 334 | 19 | 331 | Alpha-L-fucosidase. O-Glycosyl hydrolases (EC 3.2.1.-) are a widespread group of enzymes that hydrolyse the glycosidic bond between two or more carbohydrates, or between a carbohydrate and a non-carbohydrate moiety. A classification system for glycosyl hydrolases, based on sequence similarity, has led to the definition of 85 different families. This classification is available on the CAZy (CArbohydrate-Active EnZymes) web site. Because the fold of proteins is better conserved than their sequences, some of the families can be grouped in 'clans'. Family 29 encompasses alpha-L-fucosidases, which is a lysosomal enzyme responsible for hydrolyzing the alpha-1,6-linked fucose joined to the reducing-end N-acetylglucosamine of the carbohydrate moieties of glycoproteins. Deficiency of alpha-L-fucosidase results in the lysosomal storage disease fucosidosis. |
| COG3669 | AfuC | 6.61e-35 | 15 | 335 | 1 | 314 | Alpha-L-fucosidase [Carbohydrate transport and metabolism]. |
| pfam14871 | GHL6 | 1.54e-09 | 62 | 140 | 1 | 89 | Hypothetical glycosyl hydrolase 6. GHL6 is a family of hypothetical glycoside hydrolases. |
| cd07431 | PHP_PolIIIA | 0.002 | 64 | 127 | 19 | 62 | Polymerase and Histidinol Phosphatase domain of alpha-subunit of bacterial polymerase III. PolIIIAs that contain an N-terminal PHP domain have been classified into four basic groups based on genome composition, phylogenetic, and domain structural analysis: polC, dnaE1, dnaE2, and dnaE3. The PHP (also called histidinol phosphatase-2/HIS2) domain is associated with several types of DNA polymerases, such as PolIIIA and family X DNA polymerases, stand alone histidinol phosphate phosphatases (HisPPases), and a number of uncharacterized protein families. DNA polymerase III holoenzyme is one of the five eubacterial DNA polymerases that is responsible for the replication of the DNA duplex. The alpha subunit of DNA polymerase III core enzyme catalyzes the reaction for polymerizing both DNA strands. The PolIIIA PHP domain has four conserved sequence motifs and contains an invariant histidine that is involved in metal ion coordination, and like other PHP structures, exhibits a distorted (beta/alpha) 7 barrel and coordinates up to 3 metals. Initially, it was proposed that PHP region might be involved in pyrophosphate hydrolysis, but such activity has not been found. It has been shown that the PHP domain of PolIIIA has a trinuclear metal complex and is capable of proofreading activity. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| ALB28421.1 | 3.43e-206 | 3 | 332 | 6 | 337 |
| AYE37750.1 | 3.02e-205 | 3 | 332 | 6 | 337 |
| AKU60913.1 | 3.38e-205 | 4 | 335 | 7 | 341 |
| ARE44411.1 | 3.38e-205 | 4 | 335 | 7 | 341 |
| QDR74560.1 | 3.38e-205 | 4 | 335 | 7 | 341 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 6O18_A | 1.05e-205 | 4 | 335 | 7 | 341 | ChainA, AlfC [Lacticaseibacillus casei],6O18_B Chain B, AlfC [Lacticaseibacillus casei],6O18_C Chain C, AlfC [Lacticaseibacillus casei],6O18_D Chain D, AlfC [Lacticaseibacillus casei],6O1A_A Chain A, AlfC [Lacticaseibacillus casei],6O1A_B Chain B, AlfC [Lacticaseibacillus casei],6O1A_C Chain C, AlfC [Lacticaseibacillus casei],6O1A_D Chain D, AlfC [Lacticaseibacillus casei] |
| 6O1I_A | 8.61e-205 | 4 | 335 | 7 | 341 | ChainA, AlfC [Lacticaseibacillus casei],6O1I_B Chain B, AlfC [Lacticaseibacillus casei],6O1I_C Chain C, AlfC [Lacticaseibacillus casei],6O1I_D Chain D, AlfC [Lacticaseibacillus casei] |
| 6O1J_A | 1.74e-204 | 4 | 335 | 7 | 341 | ChainA, AlfC [Lacticaseibacillus casei],6O1J_B Chain B, AlfC [Lacticaseibacillus casei],6O1J_C Chain C, AlfC [Lacticaseibacillus casei],6O1J_D Chain D, AlfC [Lacticaseibacillus casei],6O1J_E Chain E, AlfC [Lacticaseibacillus casei],6O1J_F Chain F, AlfC [Lacticaseibacillus casei],6O1J_G Chain G, AlfC [Lacticaseibacillus casei],6O1J_H Chain H, AlfC [Lacticaseibacillus casei] |
| 6O1C_A | 1.74e-204 | 4 | 335 | 7 | 341 | ChainA, AlfC [Lacticaseibacillus casei],6O1C_B Chain B, AlfC [Lacticaseibacillus casei],6O1C_C Chain C, AlfC [Lacticaseibacillus casei],6O1C_D Chain D, AlfC [Lacticaseibacillus casei],6O1C_E Chain E, AlfC [Lacticaseibacillus casei],6O1C_F Chain F, AlfC [Lacticaseibacillus casei],6O1C_G Chain G, AlfC [Lacticaseibacillus casei],6O1C_H Chain H, AlfC [Lacticaseibacillus casei],6OHE_A Chain A, AlfC [Lacticaseibacillus casei],6OHE_B Chain B, AlfC [Lacticaseibacillus casei],6OHE_C Chain C, AlfC [Lacticaseibacillus casei],6OHE_D Chain D, AlfC [Lacticaseibacillus casei] |
| 7DB5_A | 1.37e-65 | 1 | 319 | 34 | 338 | ChainA, Alpha-L-fucosidase [Vibrio sp. EJY3] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| P17164 | 3.97e-23 | 1 | 331 | 45 | 356 | Tissue alpha-L-fucosidase OS=Rattus norvegicus OX=10116 GN=Fuca1 PE=1 SV=1 |
| P49713 | 1.54e-22 | 1 | 326 | 30 | 335 | Putative alpha-L-fucosidase OS=Caenorhabditis elegans OX=6239 GN=W03G11.3 PE=3 SV=2 |
| Q2KIM0 | 8.99e-22 | 1 | 331 | 52 | 363 | Tissue alpha-L-fucosidase OS=Bos taurus OX=9913 GN=FUCA1 PE=2 SV=1 |
| Q60HF8 | 1.66e-21 | 1 | 331 | 51 | 362 | Tissue alpha-L-fucosidase OS=Macaca fascicularis OX=9541 GN=FUCA1 PE=2 SV=1 |
| Q99LJ1 | 2.05e-21 | 1 | 331 | 35 | 346 | Tissue alpha-L-fucosidase OS=Mus musculus OX=10090 GN=Fuca1 PE=1 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as OTHER
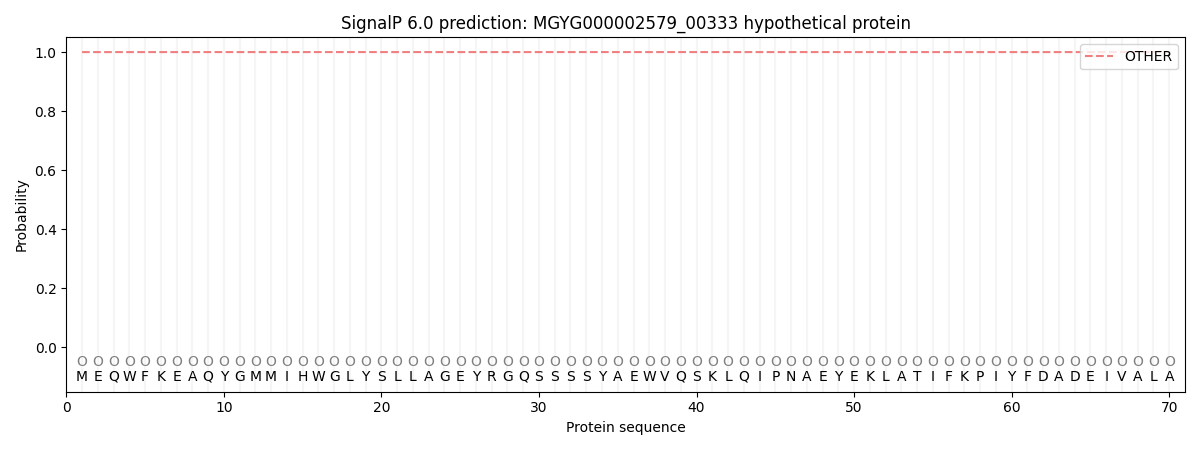
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 1.000073 | 0.000003 | 0.000000 | 0.000000 | 0.000000 | 0.000000 |
