You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000002652_00566
You are here: Home > Sequence: MGYG000002652_00566
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
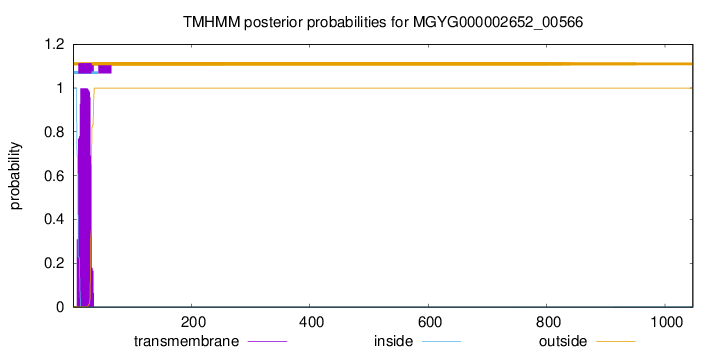
TMHMM annotations
Basic Information help
| Species | UBA4248 sp004554395 | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Myxococcota; Bradymonadia; UBA4248; UBA4248; UBA4248; UBA4248 sp004554395 | |||||||||||
| CAZyme ID | MGYG000002652_00566 | |||||||||||
| CAZy Family | GT51 | |||||||||||
| CAZyme Description | Monofunctional biosynthetic peptidoglycan transglycosylase | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 2680; End: 5823 Strand: + | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GT51 | 746 | 896 | 9.6e-47 | 0.8361581920903954 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| pfam00912 | Transgly | 3.00e-52 | 749 | 889 | 15 | 153 | Transglycosylase. The penicillin-binding proteins are bifunctional proteins consisting of transglycosylase and transpeptidase in the N- and C-terminus respectively. The transglycosylase domain catalyzes the polymerization of murein glycan chains. |
| PRK00056 | mtgA | 5.02e-46 | 748 | 891 | 61 | 202 | monofunctional biosynthetic peptidoglycan transglycosylase; Provisional |
| COG0744 | MrcB | 1.51e-45 | 700 | 918 | 31 | 242 | Membrane carboxypeptidase (penicillin-binding protein) [Cell wall/membrane/envelope biogenesis]. |
| COG5009 | MrcA | 4.50e-31 | 750 | 888 | 70 | 205 | Membrane carboxypeptidase/penicillin-binding protein [Cell wall/membrane/envelope biogenesis]. |
| COG4953 | PbpC | 6.67e-21 | 738 | 949 | 52 | 248 | Membrane carboxypeptidase/penicillin-binding protein PbpC [Cell wall/membrane/envelope biogenesis]. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| QSW91504.1 | 2.00e-47 | 542 | 919 | 257 | 607 |
| QSB27480.1 | 4.98e-47 | 540 | 919 | 255 | 607 |
| AYN05966.1 | 5.50e-45 | 542 | 919 | 257 | 607 |
| QLK35892.1 | 9.54e-45 | 555 | 891 | 270 | 577 |
| AKC23103.1 | 9.54e-45 | 555 | 891 | 270 | 577 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 3NB6_A | 7.37e-22 | 749 | 891 | 22 | 162 | Crystalstructure of Aquifex aeolicus peptidoglycan glycosyltransferase in complex with Methylphosphoryl Neryl Moenomycin [Aquifex aeolicus] |
| 2OQO_A | 1.85e-21 | 749 | 891 | 22 | 162 | Crystalstructure of a peptidoglycan glycosyltransferase from a class A PBP: insight into bacterial cell wall synthesis [Aquifex aeolicus VF5],3D3H_A Crystal structure of a complex of the peptidoglycan glycosyltransferase domain from Aquifex aeolicus and neryl moenomycin A [Aquifex aeolicus],3NB7_A Crystal structure of Aquifex Aeolicus Peptidoglycan Glycosyltransferase in complex with Decarboxylated Neryl Moenomycin [Aquifex aeolicus] |
| 5U2G_A | 4.55e-16 | 754 | 889 | 47 | 180 | 2.6Angstrom Resolution Crystal Structure of Penicillin-Binding Protein 1A from Haemophilus influenzae [Haemophilus influenzae Rd KW20],5U2G_B 2.6 Angstrom Resolution Crystal Structure of Penicillin-Binding Protein 1A from Haemophilus influenzae [Haemophilus influenzae Rd KW20] |
| 7U4H_A | 1.90e-15 | 750 | 896 | 43 | 185 | ChainA, Penicillin-binding protein 1A (Pbp1a) [Chlamydia trachomatis D/UW-3/CX],7U4H_B Chain B, Penicillin-binding protein 1A (Pbp1a) [Chlamydia trachomatis D/UW-3/CX] |
| 3DWK_A | 7.10e-15 | 754 | 895 | 34 | 173 | ChainA, Penicillin-binding protein 2 [Staphylococcus aureus subsp. aureus COL],3DWK_B Chain B, Penicillin-binding protein 2 [Staphylococcus aureus subsp. aureus COL],3DWK_C Chain C, Penicillin-binding protein 2 [Staphylococcus aureus subsp. aureus COL],3DWK_D Chain D, Penicillin-binding protein 2 [Staphylococcus aureus subsp. aureus COL] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| Q0AIQ0 | 1.22e-28 | 739 | 903 | 67 | 229 | Biosynthetic peptidoglycan transglycosylase OS=Nitrosomonas eutropha (strain DSM 101675 / C91 / Nm57) OX=335283 GN=mtgA PE=3 SV=1 |
| A3M3C7 | 1.62e-27 | 740 | 890 | 48 | 196 | Biosynthetic peptidoglycan transglycosylase OS=Acinetobacter baumannii (strain ATCC 17978 / CIP 53.77 / LMG 1025 / NCDC KC755 / 5377) OX=400667 GN=mtgA PE=3 SV=2 |
| Q7NS41 | 1.75e-27 | 737 | 903 | 49 | 213 | Biosynthetic peptidoglycan transglycosylase OS=Chromobacterium violaceum (strain ATCC 12472 / DSM 30191 / JCM 1249 / NBRC 12614 / NCIMB 9131 / NCTC 9757) OX=243365 GN=mtgA PE=3 SV=1 |
| B2HVP6 | 2.20e-27 | 740 | 890 | 48 | 196 | Biosynthetic peptidoglycan transglycosylase OS=Acinetobacter baumannii (strain ACICU) OX=405416 GN=mtgA PE=3 SV=1 |
| B0VSQ2 | 2.99e-27 | 740 | 890 | 48 | 196 | Biosynthetic peptidoglycan transglycosylase OS=Acinetobacter baumannii (strain SDF) OX=509170 GN=mtgA PE=3 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as OTHER

| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 0.984012 | 0.000350 | 0.000262 | 0.000003 | 0.000002 | 0.015408 |