You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000002653_00187
You are here: Home > Sequence: MGYG000002653_00187
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Prevotella sp002251295 | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Bacteroidota; Bacteroidia; Bacteroidales; Bacteroidaceae; Prevotella; Prevotella sp002251295 | |||||||||||
| CAZyme ID | MGYG000002653_00187 | |||||||||||
| CAZy Family | GH10 | |||||||||||
| CAZyme Description | Endo-1,4-beta-xylanase A | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 87929; End: 89152 Strand: + | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH10 | 26 | 383 | 1.6e-87 | 0.9900990099009901 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| pfam00331 | Glyco_hydro_10 | 1.90e-93 | 26 | 383 | 1 | 310 | Glycosyl hydrolase family 10. |
| smart00633 | Glyco_10 | 3.20e-83 | 85 | 381 | 1 | 263 | Glycosyl hydrolase family 10. |
| COG3693 | XynA | 1.72e-72 | 46 | 387 | 29 | 343 | Endo-1,4-beta-xylanase, GH35 family [Carbohydrate transport and metabolism]. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| QGT71449.1 | 5.66e-117 | 26 | 382 | 30 | 369 |
| QDH57038.1 | 1.14e-116 | 26 | 382 | 30 | 369 |
| SCV10721.1 | 1.48e-116 | 26 | 382 | 17 | 356 |
| QUT80664.1 | 2.28e-116 | 26 | 382 | 30 | 369 |
| QRQ57929.1 | 2.28e-116 | 26 | 382 | 30 | 369 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 2CNC_A | 2.10e-81 | 21 | 388 | 31 | 381 | Family10 xylanase [Cellvibrio mixtus] |
| 1UQY_A | 4.66e-81 | 21 | 388 | 22 | 372 | XylanaseXyn10B mutant (E262S) from Cellvibrio mixtus in complex with xylopentaose [Cellvibrio mixtus],1UQZ_A Xylanase Xyn10B mutant (E262S) from Cellvibrio mixtus in complex with 4-O-methyl glucuronic acid [Cellvibrio mixtus],1UR1_A Xylanase Xyn10B mutant (E262S) from Cellvibrio mixtus in complex with arabinofuranose alpha-1,3 linked to xylobiose [Cellvibrio mixtus],1UR2_A Xylanase Xyn10B mutant (E262S) from Cellvibrio mixtus in complex with arabinofuranose alpha 1,3 linked to xylotriose [Cellvibrio mixtus] |
| 4PMU_A | 1.31e-74 | 26 | 384 | 3 | 352 | Crystalstructure of a novel reducing-end xylose-releasing exo-oligoxylanase (XynA) belonging to GH10 family (space group P1211) [Xanthomonas citri pv. citri str. 306],4PMU_B Crystal structure of a novel reducing-end xylose-releasing exo-oligoxylanase (XynA) belonging to GH10 family (space group P1211) [Xanthomonas citri pv. citri str. 306],4PMU_C Crystal structure of a novel reducing-end xylose-releasing exo-oligoxylanase (XynA) belonging to GH10 family (space group P1211) [Xanthomonas citri pv. citri str. 306],4PMU_D Crystal structure of a novel reducing-end xylose-releasing exo-oligoxylanase (XynA) belonging to GH10 family (space group P1211) [Xanthomonas citri pv. citri str. 306],4PMU_E Crystal structure of a novel reducing-end xylose-releasing exo-oligoxylanase (XynA) belonging to GH10 family (space group P1211) [Xanthomonas citri pv. citri str. 306],4PMU_F Crystal structure of a novel reducing-end xylose-releasing exo-oligoxylanase (XynA) belonging to GH10 family (space group P1211) [Xanthomonas citri pv. citri str. 306] |
| 4PMV_A | 1.34e-74 | 26 | 384 | 4 | 353 | Crystalstructure of a novel reducing-end xylose-releasing exo-oligoxylanase (XynA) belonging to GH10 family (space group P43212) [Xanthomonas citri pv. citri str. 306],4PMV_B Crystal structure of a novel reducing-end xylose-releasing exo-oligoxylanase (XynA) belonging to GH10 family (space group P43212) [Xanthomonas citri pv. citri str. 306] |
| 6FHE_A | 8.52e-71 | 26 | 382 | 13 | 339 | Highlyactive enzymes by automated modular backbone assembly and sequence design [synthetic construct] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| P49942 | 4.55e-117 | 26 | 382 | 30 | 369 | Endo-1,4-beta-xylanase A OS=Bacteroides ovatus OX=28116 GN=xylI PE=2 SV=1 |
| P48789 | 3.36e-96 | 16 | 384 | 16 | 367 | Endo-1,4-beta-xylanase A OS=Prevotella ruminicola OX=839 GN=xynA PE=3 SV=1 |
| O69231 | 1.92e-63 | 70 | 382 | 32 | 329 | Endo-1,4-beta-xylanase B OS=Paenibacillus barcinonensis OX=198119 GN=xynB PE=1 SV=1 |
| P45703 | 2.80e-62 | 71 | 384 | 33 | 330 | Endo-1,4-beta-xylanase OS=Geobacillus stearothermophilus OX=1422 GN=xynA PE=1 SV=1 |
| P23556 | 2.37e-60 | 24 | 382 | 15 | 340 | Endo-1,4-beta-xylanase A OS=Caldicellulosiruptor saccharolyticus OX=44001 GN=xynA PE=1 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as SP
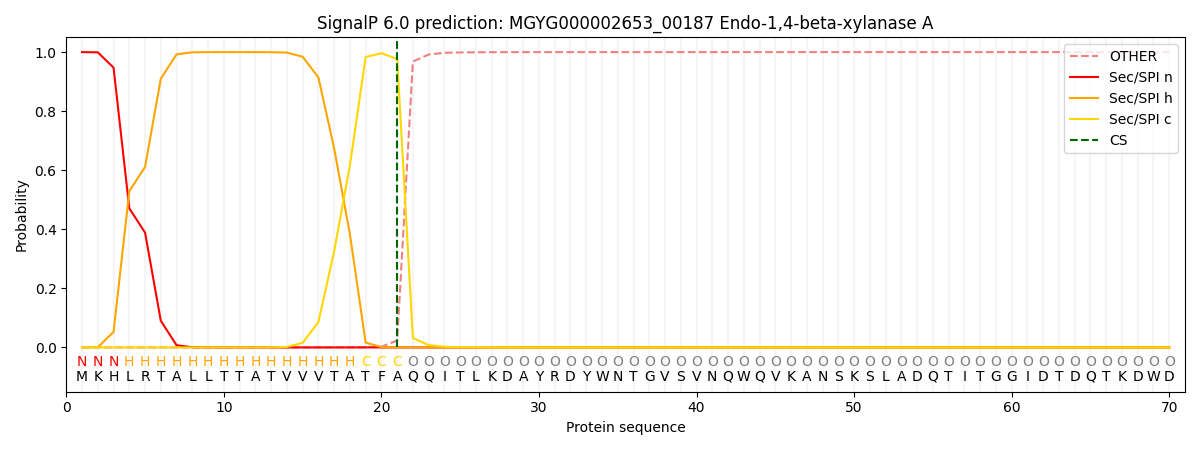
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 0.000617 | 0.998071 | 0.000457 | 0.000275 | 0.000270 | 0.000260 |
