You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000002685_00263
You are here: Home > Sequence: MGYG000002685_00263
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Firmicutes_A; Clostridia; Monoglobales_A; UBA9506; ; | |||||||||||
| CAZyme ID | MGYG000002685_00263 | |||||||||||
| CAZy Family | PL35 | |||||||||||
| CAZyme Description | hypothetical protein | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 14211; End: 18161 Strand: - | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| PL35 | 842 | 1011 | 8.1e-38 | 0.9497206703910615 |
| CBM32 | 1189 | 1308 | 6.4e-21 | 0.9112903225806451 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| pfam00754 | F5_F8_type_C | 2.01e-16 | 1185 | 1307 | 1 | 126 | F5/8 type C domain. This domain is also known as the discoidin (DS) domain family. |
| pfam16332 | DUF4962 | 4.46e-06 | 460 | 623 | 117 | 289 | Domain of unknown function (DUF4962). This family consists of uncharacterized proteins around 870 residues in length and is mainly found in various Bacteroides species. The function of this protein is unknown. |
| pfam07940 | Hepar_II_III | 0.003 | 844 | 1007 | 30 | 189 | Heparinase II/III-like protein. This family features sequences that are similar to a region of the Flavobacterium heparinum proteins heparinase II and heparinase III. The former is known to degrade heparin and heparin sulphate, whereas the latter predominantly degrades heparin sulphate. Both are secreted into the periplasmic space upon induction with heparin. |
| pfam17132 | Glyco_hydro_106 | 0.010 | 1156 | 1295 | 149 | 286 | alpha-L-rhamnosidase. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| AZS17848.1 | 5.52e-126 | 411 | 1312 | 347 | 1278 |
| ALS27107.1 | 1.81e-122 | 464 | 1311 | 678 | 1511 |
| QUH30172.1 | 3.47e-117 | 483 | 1312 | 70 | 894 |
| QNK58643.1 | 1.58e-112 | 464 | 1311 | 773 | 1609 |
| QOT12851.1 | 2.37e-100 | 462 | 1311 | 456 | 1297 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 7D29_A | 7.92e-17 | 1198 | 1314 | 24 | 135 | CBM32of AlyQ [Persicobacter sp. CCB-QB2],7D2A_A CBM32 of AlyQ in complex with 4,5-unsaturated mannuronic acid [Persicobacter sp. CCB-QB2] |
| 5XNR_A | 5.07e-15 | 1198 | 1314 | 24 | 135 | TruncatedAlyQ with CBM32 and alginate lyase domains [Persicobacter sp. CCB-QB2] |
| 2FUQ_A | 1.41e-11 | 486 | 613 | 23 | 148 | ChainA, heparinase II protein [Pedobacter heparinus],2FUQ_B Chain B, heparinase II protein [Pedobacter heparinus] |
| 3E7J_A | 1.41e-11 | 486 | 613 | 25 | 150 | ChainA, Heparinase II protein [Pedobacter heparinus],3E7J_B Chain B, Heparinase II protein [Pedobacter heparinus] |
| 3E80_A | 1.41e-11 | 486 | 613 | 25 | 150 | Structureof Heparinase II complexed with heparan sulfate degradation disaccharide product [Pedobacter heparinus],3E80_B Structure of Heparinase II complexed with heparan sulfate degradation disaccharide product [Pedobacter heparinus],3E80_C Structure of Heparinase II complexed with heparan sulfate degradation disaccharide product [Pedobacter heparinus] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| C6XZB6 | 7.89e-11 | 486 | 613 | 48 | 173 | Heparin and heparin-sulfate lyase OS=Pedobacter heparinus (strain ATCC 13125 / DSM 2366 / CIP 104194 / JCM 7457 / NBRC 12017 / NCIMB 9290 / NRRL B-14731 / HIM 762-3) OX=485917 GN=hepB PE=1 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as SP
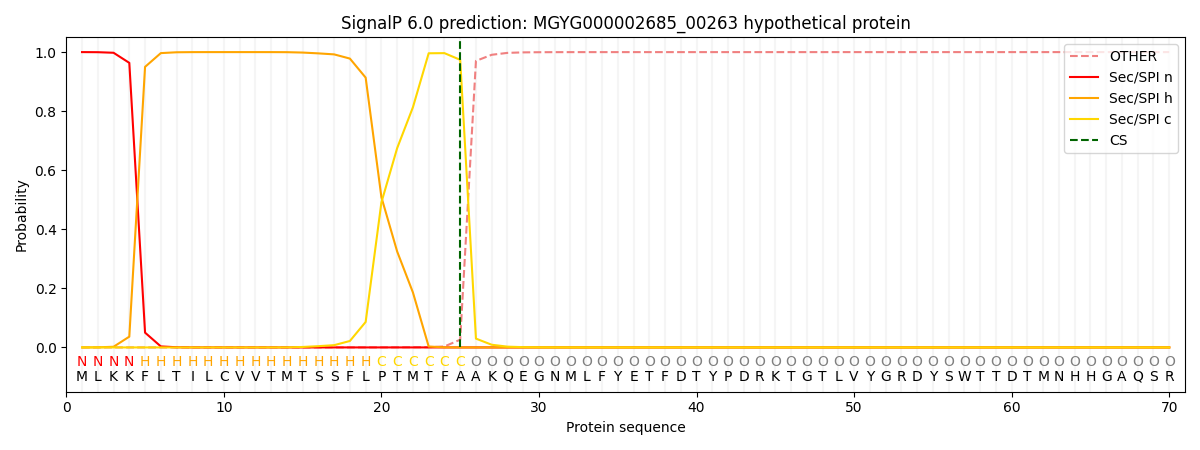
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 0.000268 | 0.998986 | 0.000203 | 0.000196 | 0.000170 | 0.000149 |