You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000002713_00778
You are here: Home > Sequence: MGYG000002713_00778
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Firmicutes_A; Clostridia; Monoglobales_A; UBA1381; UBA4716; | |||||||||||
| CAZyme ID | MGYG000002713_00778 | |||||||||||
| CAZy Family | GH5 | |||||||||||
| CAZyme Description | hypothetical protein | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 10784; End: 12697 Strand: - | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH5 | 276 | 549 | 3.9e-65 | 0.9923954372623575 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| pfam00150 | Cellulase | 4.73e-23 | 292 | 548 | 20 | 268 | Cellulase (glycosyl hydrolase family 5). |
| COG2730 | BglC | 9.68e-16 | 280 | 422 | 57 | 194 | Aryl-phospho-beta-D-glucosidase BglC, GH1 family [Carbohydrate transport and metabolism]. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| BCJ98812.1 | 1.43e-109 | 219 | 549 | 5 | 318 |
| AQR93159.1 | 6.94e-98 | 212 | 591 | 58 | 420 |
| AVM42141.1 | 5.71e-97 | 233 | 551 | 3 | 304 |
| BCJ97694.1 | 6.76e-72 | 226 | 612 | 61 | 436 |
| QBC26421.1 | 3.13e-60 | 212 | 590 | 35 | 391 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 1EQP_A | 3.04e-17 | 232 | 420 | 9 | 188 | Exo-b-(1,3)-glucanaseFrom Candida Albicans [Candida albicans] |
| 1CZ1_A | 5.43e-17 | 232 | 420 | 9 | 188 | Exo-b-(1,3)-glucanaseFrom Candida Albicans At 1.85 A Resolution [Candida albicans],1EQC_A Exo-b-(1,3)-glucanase From Candida Albicans In Complex With Castanospermine At 1.85 A [Candida albicans] |
| 4M80_A | 5.65e-17 | 232 | 420 | 14 | 193 | Thestructure of E292S glycosynthase variant of exo-1,3-beta-glucanase from Candida albicans at 1.85A resolution [Candida albicans SC5314],4M81_A The structure of E292S glycosynthase variant of exo-1,3-beta-glucanase from Candida albicans complexed with 1-fluoro-alpha-D-glucopyranoside (donor) and p-nitrophenyl beta-D-glucopyranoside (acceptor) at 1.86A resolution [Candida albicans SC5314],4M82_A The structure of E292S glycosynthase variant of exo-1,3-beta-glucanase from Candida albicans complexed with p-nitrophenyl-gentiobioside (product) at 1.6A resolution [Candida albicans SC5314] |
| 3O6A_A | 5.65e-17 | 232 | 420 | 14 | 193 | F144Y/F258YDouble Mutant of Exo-beta-1,3-glucanase from Candida albicans at 2 A [Candida albicans] |
| 3N9K_A | 5.65e-17 | 232 | 420 | 14 | 193 | F229A/E292SDouble Mutant of Exo-beta-1,3-glucanase from Candida albicans in Complex with Laminaritriose at 1.7 A [Candida albicans] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| W8QRE4 | 1.35e-30 | 221 | 549 | 10 | 338 | Beta-xylosidase OS=Phanerodontia chrysosporium OX=2822231 GN=Xyl5 PE=1 SV=2 |
| Q96V64 | 1.92e-21 | 226 | 440 | 31 | 229 | Glucan 1,3-beta-glucosidase OS=Blumeria graminis OX=34373 PE=3 SV=1 |
| Q9URU6 | 4.13e-19 | 226 | 420 | 33 | 215 | Glucan 1,3-beta-glucosidase 1 OS=Schizosaccharomyces pombe (strain 972 / ATCC 24843) OX=284812 GN=exg1 PE=2 SV=1 |
| A2RAR6 | 4.60e-18 | 213 | 419 | 23 | 211 | Probable glucan 1,3-beta-glucosidase A OS=Aspergillus niger (strain CBS 513.88 / FGSC A1513) OX=425011 GN=exgA PE=3 SV=1 |
| Q876J2 | 1.02e-17 | 231 | 420 | 53 | 235 | Sporulation-specific glucan 1,3-beta-glucosidase OS=Saccharomyces uvarum (strain ATCC 76518 / CBS 7001 / CLIB 283 / NBRC 10550 / MCYC 623 / NCYC 2669 / NRRL Y-11845) OX=659244 GN=SPR1 PE=3 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as SP
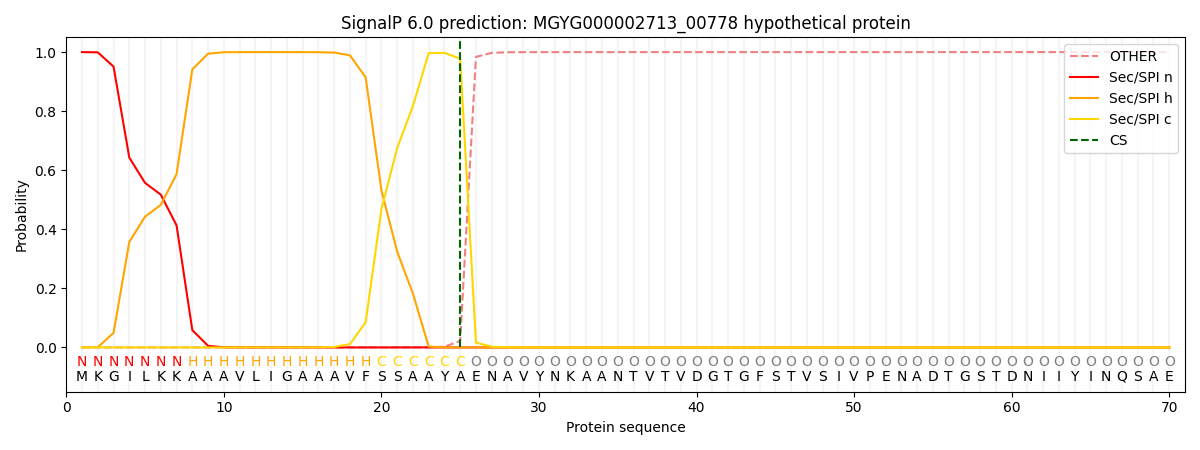
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 0.000292 | 0.998896 | 0.000226 | 0.000195 | 0.000197 | 0.000169 |
