You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000002888_00783
You are here: Home > Sequence: MGYG000002888_00783
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Corynebacterium kroppenstedtii_A | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Actinobacteriota; Actinomycetia; Mycobacteriales; Mycobacteriaceae; Corynebacterium; Corynebacterium kroppenstedtii_A | |||||||||||
| CAZyme ID | MGYG000002888_00783 | |||||||||||
| CAZy Family | CBM48 | |||||||||||
| CAZyme Description | 1,4-alpha-glucan branching enzyme GlgB | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 45982; End: 48156 Strand: + | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH13 | 271 | 570 | 4.7e-160 | 0.9933554817275747 |
| CBM48 | 120 | 210 | 9.7e-18 | 0.9078947368421053 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| PRK05402 | PRK05402 | 0.0 | 4 | 724 | 1 | 725 | 1,4-alpha-glucan branching protein GlgB. |
| PRK14706 | PRK14706 | 0.0 | 119 | 724 | 26 | 623 | glycogen branching enzyme; Provisional |
| PRK12313 | PRK12313 | 0.0 | 98 | 722 | 5 | 627 | 1,4-alpha-glucan branching protein GlgB. |
| PRK14705 | PRK14705 | 0.0 | 4 | 720 | 503 | 1220 | glycogen branching enzyme; Provisional |
| COG0296 | GlgB | 0.0 | 98 | 721 | 4 | 627 | 1,4-alpha-glucan branching enzyme [Carbohydrate transport and metabolism]. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| QRP10598.1 | 0.0 | 1 | 724 | 1 | 724 |
| ACR18012.1 | 0.0 | 1 | 724 | 1 | 724 |
| QRQ64405.1 | 0.0 | 1 | 724 | 1 | 724 |
| QRP13715.1 | 0.0 | 1 | 724 | 1 | 724 |
| ASE55676.1 | 0.0 | 4 | 724 | 11 | 731 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 3K1D_A | 0.0 | 6 | 718 | 7 | 717 | Crystalstructure of glycogen branching enzyme synonym: 1,4-alpha-D-glucan:1,4-alpha-D-GLUCAN 6-glucosyl-transferase from mycobacterium tuberculosis H37RV [Mycobacterium tuberculosis H37Rv] |
| 5GQW_A | 5.85e-225 | 3 | 723 | 24 | 774 | Crystalstructure of branching enzyme W610N mutant from Cyanothece sp. ATCC 51142 [Crocosphaera subtropica ATCC 51142],5GQX_A Crystal structure of branching enzyme W610N mutant from Cyanothece sp. ATCC 51142 in complex with maltoheptaose [Crocosphaera subtropica ATCC 51142] |
| 5GR5_A | 1.66e-224 | 3 | 723 | 24 | 774 | Crystalstructure of branching enzyme W610A mutant from Cyanothece sp. ATCC 51142 [Crocosphaera subtropica ATCC 51142] |
| 5GQU_A | 3.32e-224 | 3 | 723 | 24 | 774 | Crystalstructure of branching enzyme from Cyanothece sp. ATCC 51142 [Crocosphaera subtropica ATCC 51142],5GQV_A Crystal structure of branching enzyme from Cyanothece sp. ATCC 51142 in complex with maltohexaose [Crocosphaera subtropica ATCC 51142],5GQY_A Crystal structure of branching enzyme from Cyanothece sp. ATCC 51142 in complex with maltoheptaose [Crocosphaera subtropica ATCC 51142] |
| 5GR2_A | 4.70e-224 | 3 | 723 | 24 | 774 | Crystalstructure of branching enzyme L541A mutant from Cyanothece sp. ATCC 51142 [Crocosphaera subtropica ATCC 51142],5GR4_A Crystal structure of branching enzyme L541A mutant from Cyanothece sp. ATCC 51142 in complex with maltoheptaose [Crocosphaera subtropica ATCC 51142] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| Q1B568 | 0.0 | 5 | 721 | 22 | 740 | 1,4-alpha-glucan branching enzyme GlgB OS=Mycobacterium sp. (strain MCS) OX=164756 GN=glgB PE=3 SV=1 |
| Q5Z0W8 | 0.0 | 2 | 718 | 25 | 737 | 1,4-alpha-glucan branching enzyme GlgB OS=Nocardia farcinica (strain IFM 10152) OX=247156 GN=glgB PE=3 SV=1 |
| Q4JUK5 | 0.0 | 7 | 720 | 3 | 707 | 1,4-alpha-glucan branching enzyme GlgB OS=Corynebacterium jeikeium (strain K411) OX=306537 GN=glgB PE=3 SV=1 |
| Q6NHR6 | 0.0 | 4 | 723 | 10 | 731 | 1,4-alpha-glucan branching enzyme GlgB OS=Corynebacterium diphtheriae (strain ATCC 700971 / NCTC 13129 / Biotype gravis) OX=257309 GN=glgB PE=3 SV=1 |
| Q0SGR9 | 0.0 | 12 | 723 | 19 | 731 | 1,4-alpha-glucan branching enzyme GlgB OS=Rhodococcus jostii (strain RHA1) OX=101510 GN=glgB PE=3 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as OTHER
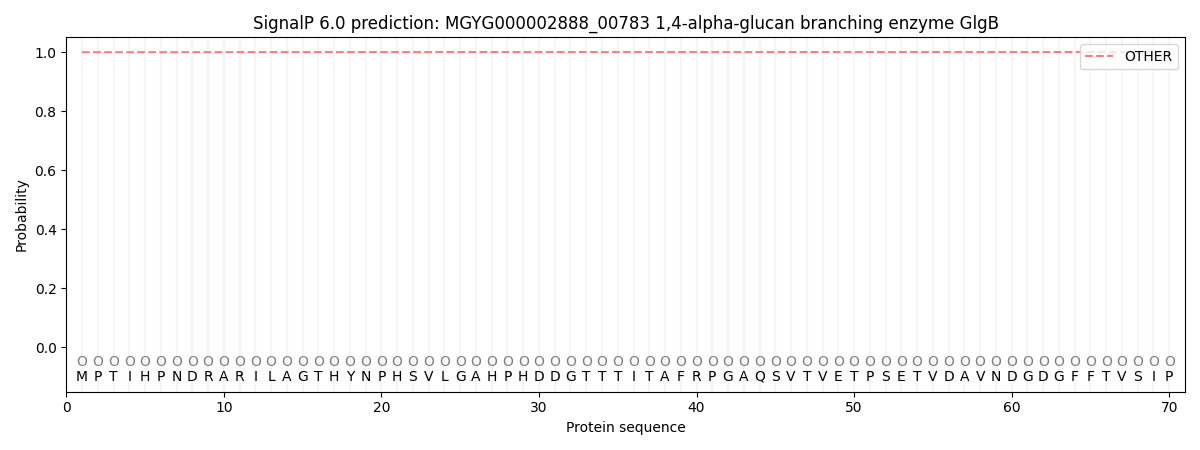
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 0.999603 | 0.000385 | 0.000049 | 0.000001 | 0.000001 | 0.000005 |
