You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000002900_01440
You are here: Home > Sequence: MGYG000002900_01440
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
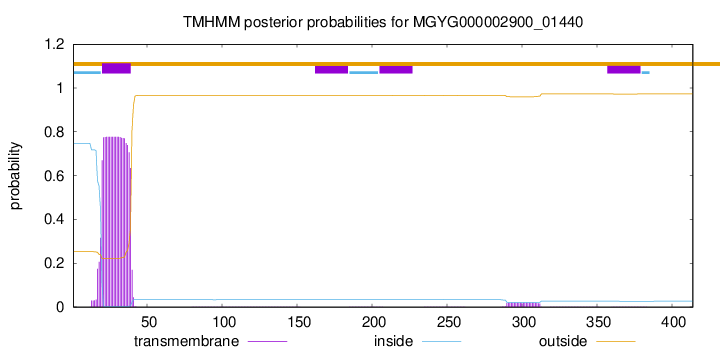
TMHMM annotations
Basic Information help
| Species | Actinomyces urogenitalis | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Actinobacteriota; Actinomycetia; Actinomycetales; Actinomycetaceae; Actinomyces; Actinomyces urogenitalis | |||||||||||
| CAZyme ID | MGYG000002900_01440 | |||||||||||
| CAZy Family | GH3 | |||||||||||
| CAZyme Description | Beta-hexosaminidase | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 7605; End: 8849 Strand: + | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH3 | 146 | 371 | 7.3e-43 | 0.9583333333333334 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| COG1472 | BglX | 7.66e-55 | 82 | 386 | 1 | 293 | Periplasmic beta-glucosidase and related glycosidases [Carbohydrate transport and metabolism]. |
| pfam00933 | Glyco_hydro_3 | 2.74e-41 | 83 | 393 | 1 | 307 | Glycosyl hydrolase family 3 N terminal domain. |
| PRK05337 | PRK05337 | 1.90e-32 | 112 | 386 | 25 | 293 | beta-hexosaminidase; Provisional |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| ALD00005.1 | 8.65e-122 | 78 | 411 | 83 | 419 |
| QKD79089.1 | 5.84e-119 | 77 | 411 | 86 | 422 |
| BDA64662.1 | 1.57e-116 | 77 | 411 | 115 | 451 |
| QQM68265.1 | 1.38e-113 | 81 | 411 | 81 | 412 |
| ARD42333.1 | 1.39e-113 | 77 | 411 | 85 | 421 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 5BU9_A | 5.25e-101 | 83 | 411 | 6 | 337 | Crystalstructure of Beta-N-acetylhexosaminidase from Beutenbergia cavernae DSM 12333 [Beutenbergia cavernae DSM 12333],5BU9_B Crystal structure of Beta-N-acetylhexosaminidase from Beutenbergia cavernae DSM 12333 [Beutenbergia cavernae DSM 12333] |
| 6K5J_A | 5.72e-27 | 83 | 411 | 12 | 337 | Structureof a glycoside hydrolase family 3 beta-N-acetylglucosaminidase from Paenibacillus sp. str. FPU-7 [Paenibacillaceae] |
| 5G1M_A | 4.14e-18 | 149 | 386 | 77 | 308 | Crystalstructure of NagZ from Pseudomonas aeruginosa [Pseudomonas aeruginosa PAO1],5G1M_B Crystal structure of NagZ from Pseudomonas aeruginosa [Pseudomonas aeruginosa PAO1],5G2M_A Crystal structure of NagZ from Pseudomonas aeruginosa in complex with N-acetylglucosamine [Pseudomonas aeruginosa PAO1],5G2M_B Crystal structure of NagZ from Pseudomonas aeruginosa in complex with N-acetylglucosamine [Pseudomonas aeruginosa PAO1],5G3R_A Crystal Structure Of Nagz From Pseudomonas Aeruginosa In Complex With N-acetylglucosamine And L-ala-1,6-anhydromurnac [Pseudomonas aeruginosa PAO1],5G3R_B Crystal Structure Of Nagz From Pseudomonas Aeruginosa In Complex With N-acetylglucosamine And L-ala-1,6-anhydromurnac [Pseudomonas aeruginosa PAO1],5G5K_A Crystal structure of NagZ from Pseudomonas aeruginosa in complex with the inhibitor 2-acetamido-1,2-dideoxynojirimycin [Pseudomonas aeruginosa PAO1],5G5K_B Crystal structure of NagZ from Pseudomonas aeruginosa in complex with the inhibitor 2-acetamido-1,2-dideoxynojirimycin [Pseudomonas aeruginosa PAO1] |
| 5G5U_A | 5.59e-18 | 149 | 386 | 77 | 308 | Crystalstructure of NagZ H174A mutant from Pseudomonas aeruginosa [Pseudomonas aeruginosa PAO1],5G5U_B Crystal structure of NagZ H174A mutant from Pseudomonas aeruginosa [Pseudomonas aeruginosa PAO1],5G6T_A Crystal structure of Zn-containing NagZ H174A mutant from Pseudomonas aeruginosa [Pseudomonas aeruginosa PAO1],5G6T_B Crystal structure of Zn-containing NagZ H174A mutant from Pseudomonas aeruginosa [Pseudomonas aeruginosa PAO1],5LY7_A Crystal structure of NagZ H174A mutant from Pseudomonas aeruginosa in complex with the inhibitor 2-acetamido-1,2-dideoxynojirimycin [Pseudomonas aeruginosa PAO1],5LY7_B Crystal structure of NagZ H174A mutant from Pseudomonas aeruginosa in complex with the inhibitor 2-acetamido-1,2-dideoxynojirimycin [Pseudomonas aeruginosa PAO1] |
| 3GS6_A | 1.66e-17 | 113 | 386 | 26 | 286 | ChainA, Beta-hexosaminidase [Vibrio cholerae],3GSM_A Chain A, Beta-hexosaminidase [Vibrio cholerae] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| Q8PMU1 | 1.71e-23 | 91 | 371 | 1 | 276 | Beta-hexosaminidase OS=Xanthomonas axonopodis pv. citri (strain 306) OX=190486 GN=nagZ PE=3 SV=1 |
| Q87BR5 | 1.74e-23 | 91 | 371 | 1 | 276 | Beta-hexosaminidase OS=Xylella fastidiosa (strain Temecula1 / ATCC 700964) OX=183190 GN=nagZ PE=3 SV=1 |
| B2I6G9 | 1.74e-23 | 91 | 371 | 1 | 276 | Beta-hexosaminidase OS=Xylella fastidiosa (strain M23) OX=405441 GN=nagZ PE=3 SV=1 |
| B2FPW9 | 2.38e-23 | 91 | 371 | 1 | 276 | Beta-hexosaminidase OS=Stenotrophomonas maltophilia (strain K279a) OX=522373 GN=nagZ PE=3 SV=1 |
| Q5H1Q0 | 3.21e-23 | 91 | 371 | 1 | 276 | Beta-hexosaminidase OS=Xanthomonas oryzae pv. oryzae (strain KACC10331 / KXO85) OX=291331 GN=nagZ PE=3 SV=2 |
SignalP and Lipop Annotations help
This protein is predicted as TAT
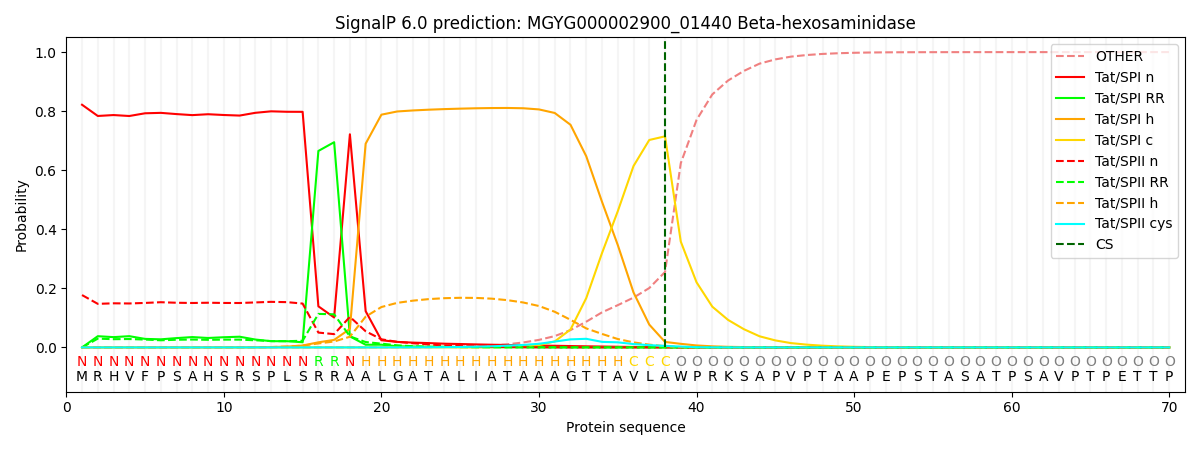
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 0.000222 | 0.000599 | 0.000084 | 0.821752 | 0.177317 | 0.000006 |