You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000002985_02017
You are here: Home > Sequence: MGYG000002985_02017
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
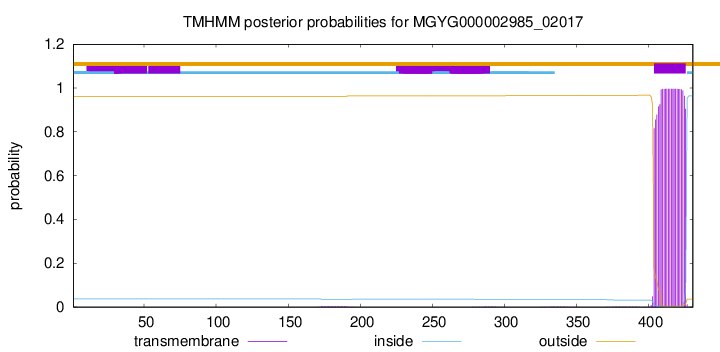
TMHMM annotations
Basic Information help
| Species | Blautia_A sp900551715 | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Firmicutes_A; Clostridia; Lachnospirales; Lachnospiraceae; Blautia_A; Blautia_A sp900551715 | |||||||||||
| CAZyme ID | MGYG000002985_02017 | |||||||||||
| CAZy Family | GH3 | |||||||||||
| CAZyme Description | hypothetical protein | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 6688; End: 7983 Strand: - | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH3 | 109 | 335 | 6.6e-35 | 0.9351851851851852 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| COG1472 | BglX | 8.32e-16 | 158 | 347 | 89 | 272 | Periplasmic beta-glucosidase and related glycosidases [Carbohydrate transport and metabolism]. |
| pfam00933 | Glyco_hydro_3 | 6.26e-12 | 157 | 347 | 93 | 308 | Glycosyl hydrolase family 3 N terminal domain. |
| PLN03080 | PLN03080 | 5.30e-08 | 138 | 346 | 94 | 343 | Probable beta-xylosidase; Provisional |
| PRK15098 | PRK15098 | 1.03e-05 | 195 | 336 | 163 | 320 | beta-glucosidase BglX. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| CBL22609.1 | 8.84e-302 | 1 | 431 | 591 | 1021 |
| QRT30388.1 | 1.79e-201 | 1 | 431 | 605 | 1035 |
| QHB24155.1 | 4.03e-200 | 1 | 431 | 605 | 1035 |
| QEI31656.1 | 4.03e-200 | 1 | 431 | 605 | 1035 |
| AMG51478.1 | 3.32e-98 | 3 | 429 | 588 | 1001 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 2X40_A | 3.33e-25 | 72 | 314 | 2 | 243 | Structureof beta-glucosidase 3B from Thermotoga neapolitana in complex with glycerol [Thermotoga neapolitana DSM 4359],2X41_A Structure of beta-glucosidase 3B from Thermotoga neapolitana in complex with glucose [Thermotoga neapolitana DSM 4359] |
| 2X42_A | 1.94e-24 | 72 | 310 | 2 | 239 | Structureof beta-glucosidase 3B from Thermotoga neapolitana in complex with alpha-D-glucose [Thermotoga neapolitana DSM 4359] |
| 3AC0_A | 2.32e-22 | 97 | 331 | 30 | 240 | Crystalstructure of Beta-glucosidase from Kluyveromyces marxianus in complex with glucose [Kluyveromyces marxianus],3AC0_B Crystal structure of Beta-glucosidase from Kluyveromyces marxianus in complex with glucose [Kluyveromyces marxianus],3AC0_C Crystal structure of Beta-glucosidase from Kluyveromyces marxianus in complex with glucose [Kluyveromyces marxianus],3AC0_D Crystal structure of Beta-glucosidase from Kluyveromyces marxianus in complex with glucose [Kluyveromyces marxianus] |
| 3ABZ_A | 4.26e-21 | 97 | 331 | 30 | 240 | Crystalstructure of Se-Met labeled Beta-glucosidase from Kluyveromyces marxianus [Kluyveromyces marxianus],3ABZ_B Crystal structure of Se-Met labeled Beta-glucosidase from Kluyveromyces marxianus [Kluyveromyces marxianus],3ABZ_C Crystal structure of Se-Met labeled Beta-glucosidase from Kluyveromyces marxianus [Kluyveromyces marxianus],3ABZ_D Crystal structure of Se-Met labeled Beta-glucosidase from Kluyveromyces marxianus [Kluyveromyces marxianus] |
| 5WUG_A | 1.46e-19 | 152 | 345 | 586 | 798 | Expression,characterization and crystal structure of a novel beta-glucosidase from Paenibacillus barengoltzii [Paenibacillus barengoltzii],5WVP_A Expression, characterization and crystal structure of a novel beta-glucosidase from Paenibacillus barengoltzii [Paenibacillus barengoltzii] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| P15885 | 4.08e-23 | 47 | 345 | 443 | 741 | Beta-glucosidase OS=Ruminococcus albus OX=1264 PE=3 SV=1 |
| P27034 | 5.20e-22 | 75 | 314 | 3 | 223 | Beta-glucosidase OS=Rhizobium radiobacter OX=358 GN=cbg-1 PE=3 SV=1 |
| P07337 | 1.70e-21 | 97 | 331 | 30 | 240 | Beta-glucosidase OS=Kluyveromyces marxianus OX=4911 PE=3 SV=1 |
| Q5AV15 | 5.47e-21 | 133 | 333 | 66 | 271 | Probable beta-glucosidase J OS=Emericella nidulans (strain FGSC A4 / ATCC 38163 / CBS 112.46 / NRRL 194 / M139) OX=227321 GN=bglJ PE=3 SV=1 |
| P16084 | 7.22e-21 | 156 | 346 | 612 | 801 | Beta-glucosidase A OS=Butyrivibrio fibrisolvens OX=831 GN=bglA PE=3 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as OTHER
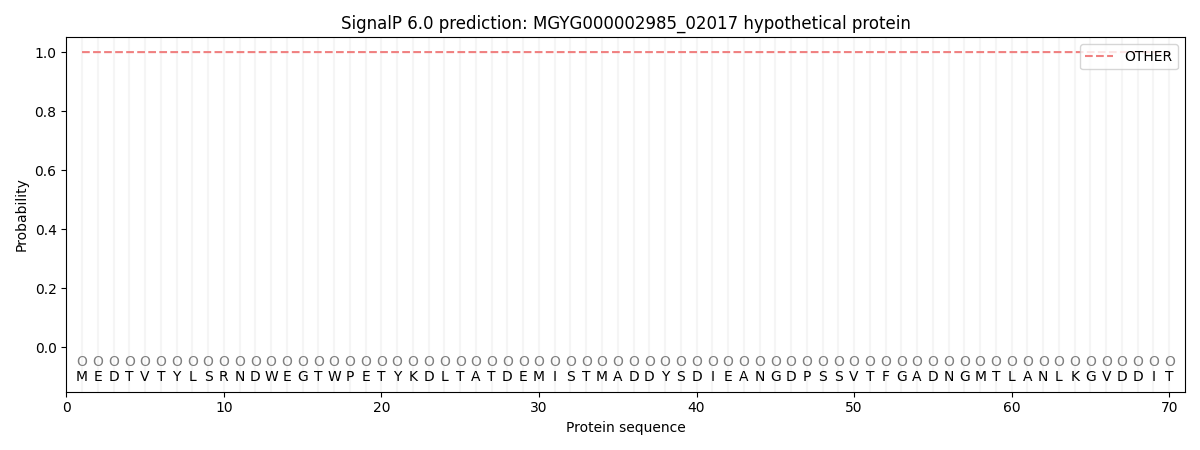
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 1.000025 | 0.000001 | 0.000000 | 0.000000 | 0.000000 | 0.000000 |