You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000003018_00601
You are here: Home > Sequence: MGYG000003018_00601
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Alistipes_A sp900549685 | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Bacteroidota; Bacteroidia; Bacteroidales; Rikenellaceae; Alistipes_A; Alistipes_A sp900549685 | |||||||||||
| CAZyme ID | MGYG000003018_00601 | |||||||||||
| CAZy Family | GH3 | |||||||||||
| CAZyme Description | Beta-glucosidase BoGH3B | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 9568; End: 11880 Strand: + | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH3 | 122 | 350 | 5.1e-78 | 0.9861111111111112 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| PRK15098 | PRK15098 | 5.47e-180 | 1 | 759 | 1 | 756 | beta-glucosidase BglX. |
| COG1472 | BglX | 7.41e-99 | 47 | 476 | 1 | 387 | Periplasmic beta-glucosidase and related glycosidases [Carbohydrate transport and metabolism]. |
| pfam00933 | Glyco_hydro_3 | 6.81e-78 | 48 | 383 | 1 | 316 | Glycosyl hydrolase family 3 N terminal domain. |
| PLN03080 | PLN03080 | 5.05e-76 | 142 | 754 | 106 | 769 | Probable beta-xylosidase; Provisional |
| pfam01915 | Glyco_hydro_3_C | 5.62e-56 | 421 | 651 | 1 | 216 | Glycosyl hydrolase family 3 C-terminal domain. This domain is involved in catalysis and may be involved in binding beta-glucan. This domain is found associated with pfam00933. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| AFL77929.1 | 4.57e-315 | 13 | 770 | 9 | 766 |
| QIU93423.1 | 5.27e-315 | 18 | 770 | 17 | 770 |
| QNL39934.1 | 4.29e-314 | 16 | 770 | 16 | 770 |
| QUT88989.1 | 3.37e-313 | 11 | 770 | 11 | 769 |
| ALJ59996.1 | 3.88e-312 | 11 | 770 | 11 | 769 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 5XXL_A | 7.46e-149 | 34 | 770 | 8 | 754 | Crystalstructure of GH3 beta-glucosidase from Bacteroides thetaiotaomicron [Bacteroides thetaiotaomicron VPI-5482],5XXL_B Crystal structure of GH3 beta-glucosidase from Bacteroides thetaiotaomicron [Bacteroides thetaiotaomicron VPI-5482],5XXM_A Crystal structure of GH3 beta-glucosidase from Bacteroides thetaiotaomicron in complex with gluconolactone [Bacteroides thetaiotaomicron VPI-5482],5XXM_B Crystal structure of GH3 beta-glucosidase from Bacteroides thetaiotaomicron in complex with gluconolactone [Bacteroides thetaiotaomicron VPI-5482] |
| 5XXN_A | 4.14e-148 | 34 | 770 | 8 | 754 | CrystalStructure of mutant (D286N) beta-glucosidase from Bacteroides thetaiotaomicron in complex with sophorose [Bacteroides thetaiotaomicron VPI-5482],5XXN_B Crystal Structure of mutant (D286N) beta-glucosidase from Bacteroides thetaiotaomicron in complex with sophorose [Bacteroides thetaiotaomicron VPI-5482],5XXO_A Crystal structure of mutant (D286N) GH3 beta-glucosidase from Bacteroides thetaiotaomicron in complex with sophorotriose [Bacteroides thetaiotaomicron VPI-5482],5XXO_B Crystal structure of mutant (D286N) GH3 beta-glucosidase from Bacteroides thetaiotaomicron in complex with sophorotriose [Bacteroides thetaiotaomicron VPI-5482] |
| 5Z9S_A | 1.64e-143 | 29 | 769 | 17 | 787 | Functionaland Structural Characterization of a beta-Glucosidase Involved in Saponin Metabolism from Intestinal Bacteria [Bifidobacterium longum],5Z9S_B Functional and Structural Characterization of a beta-Glucosidase Involved in Saponin Metabolism from Intestinal Bacteria [Bifidobacterium longum] |
| 5Z87_A | 4.75e-143 | 119 | 769 | 113 | 784 | ChainA, EmGH1 [Aurantiacibacter marinus],5Z87_B Chain B, EmGH1 [Aurantiacibacter marinus] |
| 6R5I_A | 1.73e-138 | 37 | 758 | 4 | 723 | ChainA, Periplasmic beta-glucosidase [Pseudomonas aeruginosa PAO1],6R5I_B Chain B, Periplasmic beta-glucosidase [Pseudomonas aeruginosa PAO1],6R5N_A Chain A, Periplasmic beta-glucosidase [Pseudomonas aeruginosa PAO1],6R5N_B Chain B, Periplasmic beta-glucosidase [Pseudomonas aeruginosa PAO1] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| T2KMH0 | 2.72e-155 | 115 | 770 | 51 | 723 | Beta-xylosidase OS=Formosa agariphila (strain DSM 15362 / KCTC 12365 / LMG 23005 / KMM 3901 / M-2Alg 35-1) OX=1347342 GN=BN863_22130 PE=1 SV=1 |
| Q56078 | 6.77e-128 | 40 | 758 | 38 | 755 | Periplasmic beta-glucosidase OS=Salmonella typhimurium (strain LT2 / SGSC1412 / ATCC 700720) OX=99287 GN=bglX PE=3 SV=2 |
| P33363 | 1.93e-122 | 40 | 758 | 38 | 755 | Periplasmic beta-glucosidase OS=Escherichia coli (strain K12) OX=83333 GN=bglX PE=3 SV=2 |
| T2KMH9 | 6.78e-109 | 29 | 759 | 29 | 746 | Putative beta-xylosidase OS=Formosa agariphila (strain DSM 15362 / KCTC 12365 / LMG 23005 / KMM 3901 / M-2Alg 35-1) OX=1347342 GN=BN863_22230 PE=2 SV=1 |
| Q23892 | 2.37e-107 | 40 | 755 | 80 | 813 | Lysosomal beta glucosidase OS=Dictyostelium discoideum OX=44689 GN=gluA PE=1 SV=2 |
SignalP and Lipop Annotations help
This protein is predicted as LIPO
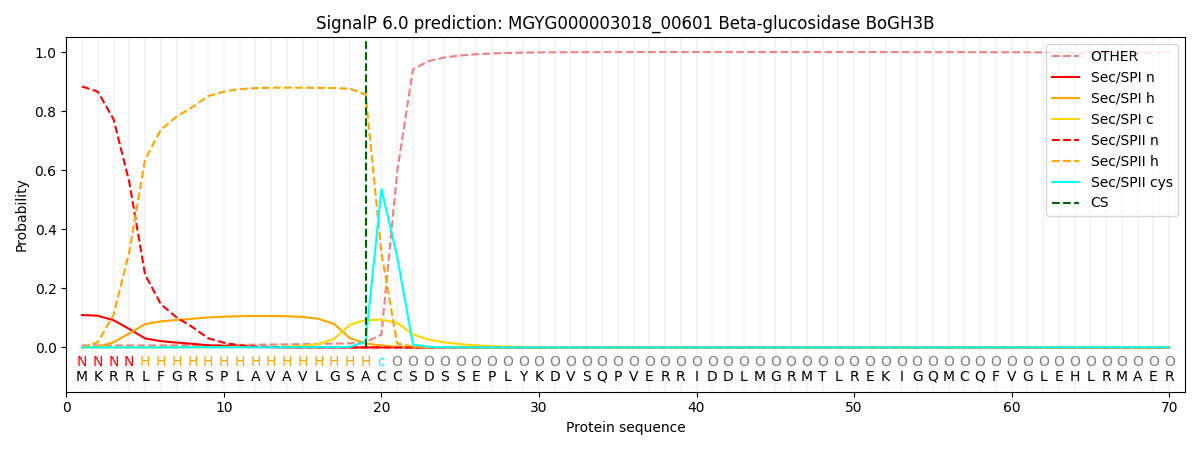
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 0.008683 | 0.103761 | 0.886429 | 0.000312 | 0.000517 | 0.000303 |
