You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000003072_02487
You are here: Home > Sequence: MGYG000003072_02487
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
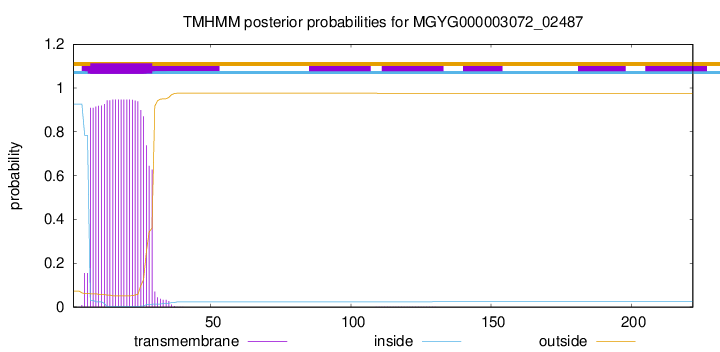
TMHMM annotations
Basic Information help
| Species | Paenibacillus amylolyticus | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Firmicutes; Bacilli; Paenibacillales; Paenibacillaceae; Paenibacillus; Paenibacillus amylolyticus | |||||||||||
| CAZyme ID | MGYG000003072_02487 | |||||||||||
| CAZy Family | PL3 | |||||||||||
| CAZyme Description | Pectate lyase A | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 11136; End: 11804 Strand: + | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| PL3 | 28 | 199 | 5.1e-76 | 0.9826589595375722 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| pfam03211 | Pectate_lyase | 5.33e-75 | 31 | 213 | 14 | 200 | Pectate lyase. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| QZN77940.1 | 8.02e-152 | 1 | 222 | 1 | 222 |
| APO44009.1 | 3.66e-148 | 1 | 222 | 1 | 222 |
| CAB40884.1 | 1.43e-145 | 1 | 222 | 1 | 222 |
| QKS60292.1 | 1.43e-145 | 1 | 222 | 1 | 222 |
| ADB78774.1 | 2.88e-145 | 1 | 222 | 1 | 222 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 1EE6_A | 1.13e-110 | 26 | 222 | 1 | 197 | CrystalStructure Of Pectate Lyase From Bacillus Sp. Strain Ksm-P15. [Bacillus sp. KSM-P15] |
| 4EW9_A | 2.74e-58 | 29 | 207 | 9 | 185 | Theliganded structure of C. bescii family 3 pectate lyase [Caldicellulosiruptor bescii DSM 6725],4EW9_B The liganded structure of C. bescii family 3 pectate lyase [Caldicellulosiruptor bescii DSM 6725] |
| 3T9G_A | 2.82e-58 | 29 | 207 | 10 | 186 | Thecrystal structure of family 3 pectate lyase from Caldicellulosiruptor bescii [Caldicellulosiruptor bescii],3T9G_B The crystal structure of family 3 pectate lyase from Caldicellulosiruptor bescii [Caldicellulosiruptor bescii] |
| 4Z03_A | 2.93e-57 | 29 | 207 | 18 | 194 | C.bescii Family 3 pectate lyase double mutant K108A in complex with trigalacturonic acid [Caldicellulosiruptor bescii DSM 6725],4Z03_B C. bescii Family 3 pectate lyase double mutant K108A in complex with trigalacturonic acid [Caldicellulosiruptor bescii DSM 6725] |
| 4Z05_A | 2.93e-57 | 29 | 207 | 18 | 194 | C.bescii Family 3 pectate lyase mutant E84A [Caldicellulosiruptor bescii DSM 6725],4Z05_B C. bescii Family 3 pectate lyase mutant E84A [Caldicellulosiruptor bescii DSM 6725] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| Q9X6Z2 | 2.86e-146 | 1 | 222 | 1 | 222 | Pectate lyase A OS=Paenibacillus barcinonensis OX=198119 GN=pelA PE=1 SV=1 |
| D3JTC1 | 5.76e-146 | 1 | 222 | 1 | 222 | Pectate lyase A OS=Paenibacillus amylolyticus OX=1451 GN=pelA PE=1 SV=1 |
| O34310 | 3.23e-70 | 1 | 202 | 1 | 203 | Pectate lyase C OS=Bacillus subtilis (strain 168) OX=224308 GN=pelC PE=1 SV=1 |
| Q65EF5 | 1.31e-69 | 1 | 202 | 1 | 203 | Pectate lyase C OS=Bacillus licheniformis (strain ATCC 14580 / DSM 13 / JCM 2505 / CCUG 7422 / NBRC 12200 / NCIMB 9375 / NCTC 10341 / NRRL NRS-1264 / Gibson 46) OX=279010 GN=pelC PE=3 SV=2 |
| A1DCY5 | 7.79e-25 | 36 | 160 | 36 | 160 | Probable pectate lyase D OS=Neosartorya fischeri (strain ATCC 1020 / DSM 3700 / CBS 544.65 / FGSC A1164 / JCM 1740 / NRRL 181 / WB 181) OX=331117 GN=plyD PE=3 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as SP
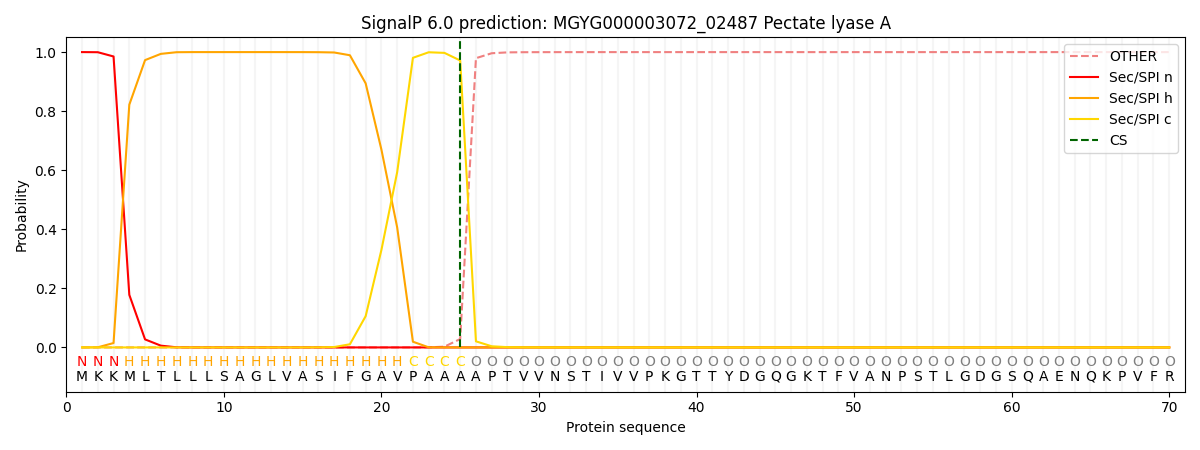
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 0.000241 | 0.999032 | 0.000188 | 0.000192 | 0.000174 | 0.000157 |