You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000003097_02234
You are here: Home > Sequence: MGYG000003097_02234
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Phocaeicola sp000436795 | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Bacteroidota; Bacteroidia; Bacteroidales; Bacteroidaceae; Phocaeicola; Phocaeicola sp000436795 | |||||||||||
| CAZyme ID | MGYG000003097_02234 | |||||||||||
| CAZy Family | PL10 | |||||||||||
| CAZyme Description | hypothetical protein | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 27399; End: 29600 Strand: - | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| PL10 | 65 | 353 | 3.6e-98 | 0.9965277777777778 |
| CE19 | 403 | 732 | 2e-29 | 0.8614457831325302 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| pfam09492 | Pec_lyase | 2.03e-95 | 65 | 354 | 1 | 289 | Pectic acid lyase. Members of this family are isozymes of pectate lyase (EC:4.2.2.2), also called polygalacturonic transeliminase and alpha-1,4-D-endopolygalacturonic acid lyase. |
| TIGR02474 | pec_lyase | 5.06e-94 | 65 | 353 | 1 | 289 | pectate lyase, PelA/Pel-15E family. Members of this family are isozymes of pectate lyase (EC 4.2.2.2), also called polygalacturonic transeliminase and alpha-1,4-D-endopolygalacturonic acid lyase. [Energy metabolism, Biosynthesis and degradation of polysaccharides] |
| COG0412 | DLH | 1.78e-20 | 442 | 619 | 1 | 142 | Dienelactone hydrolase [Secondary metabolites biosynthesis, transport and catabolism]. |
| pfam12715 | Abhydrolase_7 | 2.09e-17 | 402 | 686 | 45 | 326 | Abhydrolase family. This is a family of probable bacterial abhydrolases. |
| COG1506 | DAP2 | 1.67e-10 | 436 | 679 | 358 | 568 | Dipeptidyl aminopeptidase/acylaminoacyl peptidase [Amino acid transport and metabolism]. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| ADY37157.1 | 0.0 | 1 | 730 | 1 | 730 |
| QDU41600.1 | 2.70e-114 | 27 | 364 | 51 | 390 |
| QDT46035.1 | 1.65e-112 | 27 | 353 | 51 | 378 |
| QDU80650.1 | 8.57e-107 | 27 | 364 | 48 | 387 |
| BAV08751.1 | 2.44e-91 | 19 | 364 | 22 | 382 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 1R76_A | 5.31e-51 | 15 | 363 | 47 | 407 | ChainA, pectate lyase [Niveispirillum irakense] |
| 1GXM_A | 1.17e-37 | 58 | 345 | 37 | 314 | Family10 polysaccharide lyase from Cellvibrio cellulosa [Cellvibrio japonicus],1GXM_B Family 10 polysaccharide lyase from Cellvibrio cellulosa [Cellvibrio japonicus],1GXN_A Family 10 polysaccharide lyase from Cellvibrio cellulosa [Cellvibrio japonicus] |
| 1GXO_A | 1.39e-36 | 58 | 345 | 37 | 314 | MutantD189A of Family 10 polysaccharide lyase from Cellvibrio cellulosa in complex with trigalaturonic acid [Cellvibrio japonicus] |
| 3G8Y_A | 1.21e-13 | 383 | 720 | 27 | 352 | ChainA, SusD/RagB-associated esterase-like protein [Phocaeicola vulgatus ATCC 8482] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| O34973 | 7.69e-27 | 408 | 726 | 2 | 297 | Putative hydrolase YtaP OS=Bacillus subtilis (strain 168) OX=224308 GN=ytaP PE=3 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as SP
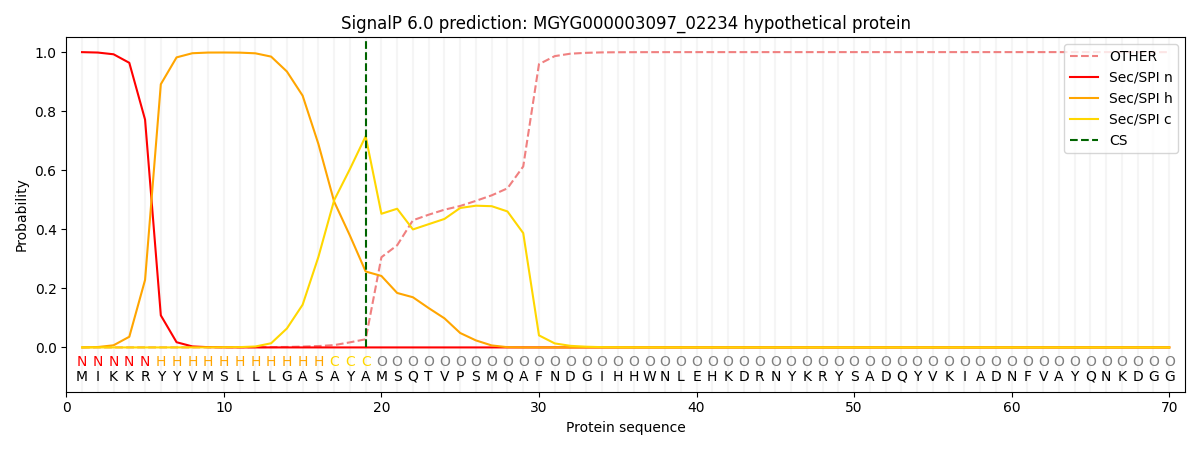
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 0.000852 | 0.998048 | 0.000278 | 0.000341 | 0.000246 | 0.000212 |
