You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000003104_00961
You are here: Home > Sequence: MGYG000003104_00961
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | UMGS1601 sp900545345 | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Firmicutes_A; Clostridia; Oscillospirales; Ruminococcaceae; UMGS1601; UMGS1601 sp900545345 | |||||||||||
| CAZyme ID | MGYG000003104_00961 | |||||||||||
| CAZy Family | GH5 | |||||||||||
| CAZyme Description | hypothetical protein | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 70038; End: 71429 Strand: + | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH5 | 80 | 353 | 3.6e-101 | 0.9891304347826086 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| pfam00150 | Cellulase | 7.23e-59 | 80 | 353 | 16 | 269 | Cellulase (glycosyl hydrolase family 5). |
| COG2730 | BglC | 5.92e-27 | 41 | 363 | 44 | 370 | Aryl-phospho-beta-D-glucosidase BglC, GH1 family [Carbohydrate transport and metabolism]. |
| smart00740 | PASTA | 6.88e-07 | 400 | 463 | 3 | 67 | PASTA domain. |
| NF033483 | PknB_PASTA_kin | 1.37e-06 | 400 | 463 | 431 | 496 | Stk1 family PASTA domain-containing Ser/Thr kinase. |
| COG2815 | PASTA | 7.77e-06 | 400 | 463 | 91 | 157 | PASTA domain, binds beta-lactams [Cell wall/membrane/envelope biogenesis]. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| CBK96026.1 | 3.44e-151 | 34 | 383 | 93 | 447 |
| CBL35063.1 | 1.55e-150 | 34 | 383 | 138 | 492 |
| CCO05659.1 | 5.39e-145 | 2 | 384 | 3 | 395 |
| CBL17363.1 | 1.71e-101 | 5 | 373 | 5 | 371 |
| QQA02033.1 | 2.77e-94 | 33 | 358 | 37 | 357 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 6GL2_A | 1.99e-77 | 34 | 372 | 7 | 335 | Structureof ZgEngAGH5_4 wild type at 1.2 Angstrom resolution [Zobellia galactanivorans] |
| 6GL0_A | 7.25e-76 | 36 | 372 | 3 | 329 | Structureof ZgEngAGH5_4 in complex with a cellotriose [Zobellia galactanivorans],6GL0_B Structure of ZgEngAGH5_4 in complex with a cellotriose [Zobellia galactanivorans],6GL0_C Structure of ZgEngAGH5_4 in complex with a cellotriose [Zobellia galactanivorans] |
| 3AYR_A | 9.82e-76 | 35 | 353 | 14 | 328 | GH5endoglucanase EglA from a ruminal fungus [Piromyces rhizinflatus] |
| 3AYS_A | 7.67e-75 | 35 | 353 | 14 | 328 | GH5endoglucanase from a ruminal fungus in complex with cellotriose [Piromyces rhizinflatus] |
| 4IM4_A | 2.87e-73 | 36 | 376 | 2 | 335 | ChainA, Endoglucanase E [Acetivibrio thermocellus],4IM4_B Chain B, Endoglucanase E [Acetivibrio thermocellus],4IM4_C Chain C, Endoglucanase E [Acetivibrio thermocellus],4IM4_D Chain D, Endoglucanase E [Acetivibrio thermocellus],4IM4_E Chain E, Endoglucanase E [Acetivibrio thermocellus],4IM4_F Chain F, Endoglucanase E [Acetivibrio thermocellus] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| P23660 | 1.81e-79 | 29 | 372 | 15 | 359 | Endoglucanase A OS=Ruminococcus albus OX=1264 GN=celA PE=1 SV=1 |
| Q12647 | 2.67e-70 | 36 | 353 | 20 | 331 | Endoglucanase B OS=Neocallimastix patriciarum OX=4758 GN=CELB PE=2 SV=1 |
| O08342 | 9.84e-68 | 11 | 353 | 10 | 369 | Endoglucanase A OS=Paenibacillus barcinonensis OX=198119 GN=celA PE=1 SV=1 |
| P10477 | 1.35e-67 | 36 | 376 | 52 | 385 | Cellulase/esterase CelE OS=Acetivibrio thermocellus (strain ATCC 27405 / DSM 1237 / JCM 9322 / NBRC 103400 / NCIMB 10682 / NRRL B-4536 / VPI 7372) OX=203119 GN=celE PE=1 SV=2 |
| P28621 | 8.39e-66 | 30 | 353 | 31 | 341 | Endoglucanase B OS=Clostridium cellulovorans (strain ATCC 35296 / DSM 3052 / OCM 3 / 743B) OX=573061 GN=engB PE=3 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as LIPO
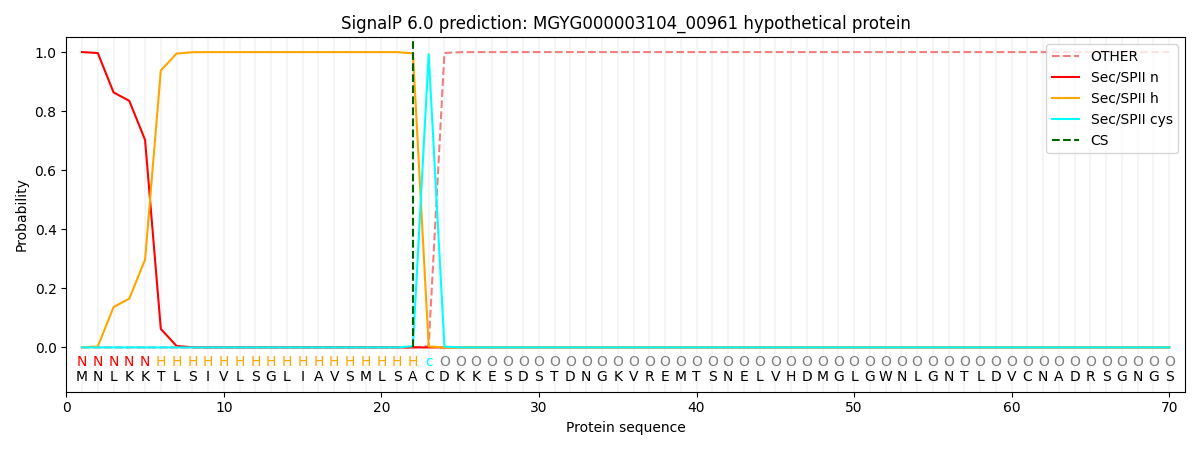
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 0.000000 | 0.000002 | 1.000064 | 0.000000 | 0.000000 | 0.000000 |
