You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000003114_01325
You are here: Home > Sequence: MGYG000003114_01325
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
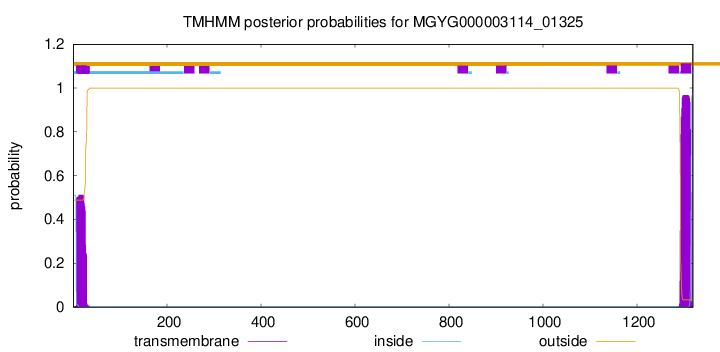
TMHMM annotations
Basic Information help
| Species | Clostridium sp900547475 | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Firmicutes_A; Clostridia; Clostridiales; Clostridiaceae; Clostridium; Clostridium sp900547475 | |||||||||||
| CAZyme ID | MGYG000003114_01325 | |||||||||||
| CAZy Family | GH29 | |||||||||||
| CAZyme Description | hypothetical protein | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 64288; End: 68247 Strand: + | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH29 | 39 | 367 | 1.6e-66 | 0.869942196531792 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| COG3669 | AfuC | 6.54e-91 | 40 | 504 | 8 | 430 | Alpha-L-fucosidase [Carbohydrate transport and metabolism]. |
| pfam01120 | Alpha_L_fucos | 8.28e-29 | 79 | 364 | 77 | 323 | Alpha-L-fucosidase. |
| smart00812 | Alpha_L_fucos | 5.13e-27 | 50 | 372 | 23 | 340 | Alpha-L-fucosidase. O-Glycosyl hydrolases (EC 3.2.1.-) are a widespread group of enzymes that hydrolyse the glycosidic bond between two or more carbohydrates, or between a carbohydrate and a non-carbohydrate moiety. A classification system for glycosyl hydrolases, based on sequence similarity, has led to the definition of 85 different families. This classification is available on the CAZy (CArbohydrate-Active EnZymes) web site. Because the fold of proteins is better conserved than their sequences, some of the families can be grouped in 'clans'. Family 29 encompasses alpha-L-fucosidases, which is a lysosomal enzyme responsible for hydrolyzing the alpha-1,6-linked fucose joined to the reducing-end N-acetylglucosamine of the carbohydrate moieties of glycoproteins. Deficiency of alpha-L-fucosidase results in the lysosomal storage disease fucosidosis. |
| smart00237 | Calx_beta | 1.57e-12 | 645 | 729 | 1 | 86 | Domains in Na-Ca exchangers and integrin-beta4. Domain in Na-Ca exchangers and integrin subunit beta4 (and some cyanobacterial proteins) |
| pfam03160 | Calx-beta | 7.38e-11 | 645 | 733 | 1 | 91 | Calx-beta domain. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| QUB39236.1 | 5.35e-218 | 39 | 760 | 161 | 856 |
| SLK21779.1 | 5.67e-216 | 39 | 763 | 42 | 752 |
| ATD55836.1 | 5.67e-216 | 39 | 763 | 42 | 752 |
| QBJ76136.1 | 5.67e-216 | 39 | 763 | 42 | 752 |
| ATD56491.1 | 5.67e-216 | 39 | 763 | 42 | 752 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 6ORG_A | 1.54e-147 | 40 | 505 | 8 | 449 | Crystalstructure of SpGH29 [Streptococcus pneumoniae TIGR4],6ORG_B Crystal structure of SpGH29 [Streptococcus pneumoniae TIGR4] |
| 6OR4_A | 2.19e-146 | 40 | 505 | 8 | 449 | Crystalstructure of SpGH29 [Streptococcus pneumoniae TIGR4],6OR4_B Crystal structure of SpGH29 [Streptococcus pneumoniae TIGR4],6ORH_A Crystal structure of SpGH29 [Streptococcus pneumoniae TIGR4],6ORH_B Crystal structure of SpGH29 [Streptococcus pneumoniae TIGR4] |
| 6ORF_A | 2.26e-146 | 40 | 505 | 8 | 449 | Crystalstructure of SpGH29 [Streptococcus pneumoniae TIGR4],6ORF_B Crystal structure of SpGH29 [Streptococcus pneumoniae TIGR4] |
| 6TR3_A | 4.63e-135 | 36 | 538 | 21 | 545 | Ruminococcusgnavus GH29 fucosidase E1_10125 in complex with fucose [[Ruminococcus] gnavus E1] |
| 6TR4_A | 6.66e-134 | 36 | 538 | 21 | 545 | Ruminococcusgnavus GH29 fucosidase E1_10125 D221A mutant in complex with fucose [[Ruminococcus] gnavus E1],6TR4_B Ruminococcus gnavus GH29 fucosidase E1_10125 D221A mutant in complex with fucose [[Ruminococcus] gnavus E1] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| Q8GW72 | 5.32e-109 | 42 | 504 | 37 | 477 | Alpha-L-fucosidase 1 OS=Arabidopsis thaliana OX=3702 GN=FUC1 PE=1 SV=2 |
| Q7XUR3 | 9.04e-104 | 35 | 498 | 32 | 470 | Putative alpha-L-fucosidase 1 OS=Oryza sativa subsp. japonica OX=39947 GN=Os04g0560400 PE=3 SV=2 |
| P10901 | 7.62e-10 | 50 | 254 | 82 | 262 | Alpha-L-fucosidase OS=Dictyostelium discoideum OX=44689 GN=alfA PE=3 SV=1 |
| E8MGH9 | 6.10e-07 | 1121 | 1240 | 1660 | 1783 | Beta-L-arabinobiosidase OS=Bifidobacterium longum subsp. longum (strain ATCC 15707 / DSM 20219 / JCM 1217 / NCTC 11818 / E194b) OX=565042 GN=hypBA2 PE=1 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as SP
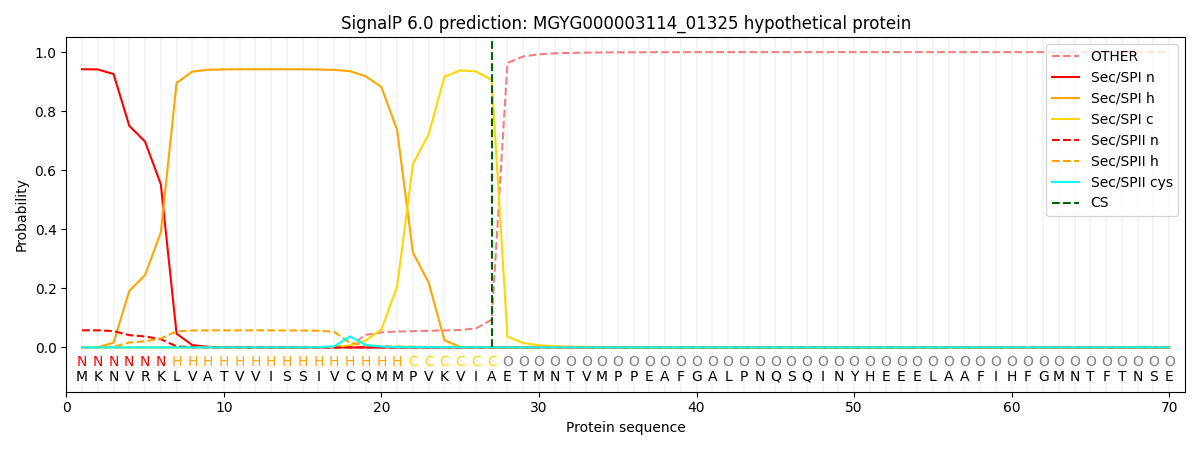
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 0.000509 | 0.937341 | 0.061382 | 0.000259 | 0.000251 | 0.000219 |