You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000003146_00388
You are here: Home > Sequence: MGYG000003146_00388
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
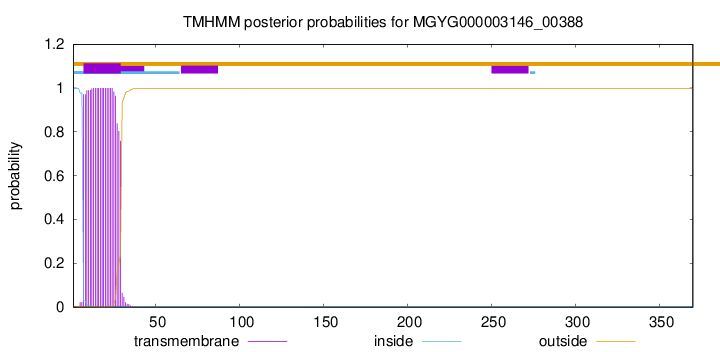
TMHMM annotations
Basic Information help
| Species | Streptococcus oralis_S | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Firmicutes; Bacilli; Lactobacillales; Streptococcaceae; Streptococcus; Streptococcus oralis_S | |||||||||||
| CAZyme ID | MGYG000003146_00388 | |||||||||||
| CAZy Family | GH73 | |||||||||||
| CAZyme Description | hypothetical protein | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 68625; End: 69737 Strand: + | |||||||||||
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| PRK08581 | PRK08581 | 3.54e-44 | 259 | 370 | 505 | 619 | amidase domain-containing protein. |
| COG3942 | COG3942 | 3.59e-38 | 189 | 370 | 1 | 173 | Surface antigen [Cell wall/membrane/envelope biogenesis]. |
| pfam05257 | CHAP | 2.02e-19 | 259 | 344 | 2 | 83 | CHAP domain. This domain corresponds to an amidase function. Many of these proteins are involved in cell wall metabolism of bacteria. This domain is found at the N-terminus of Escherichia coli gss, where it functions as a glutathionylspermidine amidase EC:3.5.1.78. This domain is found to be the catalytic domain of PlyCA. CHAP is the amidase domain of bifunctional Escherichia coli glutathionylspermidine synthetase/amidase, and it catalyzes the hydrolysis of Gsp (glutathionylspermidine) into glutathione and spermidine. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| QQY18169.1 | 8.26e-251 | 1 | 370 | 1 | 370 |
| QLL96560.1 | 1.12e-248 | 1 | 370 | 1 | 370 |
| QEW08839.1 | 1.56e-216 | 1 | 369 | 1 | 369 |
| QMI51832.1 | 2.35e-211 | 1 | 370 | 1 | 370 |
| CAD0159858.1 | 1.92e-210 | 1 | 370 | 1 | 370 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 5T1Q_A | 7.02e-32 | 262 | 370 | 248 | 359 | ChainA, N-acetylmuramoyl-L-alanine amidase domain-containing protein SAOUHSC_02979 [Staphylococcus aureus subsp. aureus NCTC 8325],5T1Q_B Chain B, N-acetylmuramoyl-L-alanine amidase domain-containing protein SAOUHSC_02979 [Staphylococcus aureus subsp. aureus NCTC 8325],5T1Q_C Chain C, N-acetylmuramoyl-L-alanine amidase domain-containing protein SAOUHSC_02979 [Staphylococcus aureus subsp. aureus NCTC 8325],5T1Q_D Chain D, N-acetylmuramoyl-L-alanine amidase domain-containing protein SAOUHSC_02979 [Staphylococcus aureus subsp. aureus NCTC 8325] |
| 2K3A_A | 8.55e-19 | 243 | 367 | 35 | 152 | ChainA, CHAP domain protein [Staphylococcus saprophyticus subsp. saprophyticus ATCC 15305 = NCTC 7292] |
| 2LRJ_A | 3.38e-16 | 262 | 367 | 11 | 111 | ChainA, Staphyloxanthin biosynthesis protein, putative [Staphylococcus aureus subsp. aureus COL] |
| 4CGK_A | 4.25e-08 | 259 | 347 | 284 | 365 | Crystalstructure of the essential protein PcsB from Streptococcus pneumoniae [Streptococcus pneumoniae D39],4CGK_B Crystal structure of the essential protein PcsB from Streptococcus pneumoniae [Streptococcus pneumoniae D39] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| Q2G222 | 1.06e-29 | 262 | 370 | 508 | 619 | N-acetylmuramoyl-L-alanine amidase domain-containing protein SAOUHSC_02979 OS=Staphylococcus aureus (strain NCTC 8325 / PS 47) OX=93061 GN=SAOUHSC_02979 PE=1 SV=1 |
| Q2YVT4 | 1.06e-18 | 184 | 367 | 151 | 334 | N-acetylmuramoyl-L-alanine amidase sle1 OS=Staphylococcus aureus (strain bovine RF122 / ET3-1) OX=273036 GN=sle1 PE=3 SV=1 |
| Q49UX4 | 1.55e-17 | 221 | 367 | 187 | 326 | N-acetylmuramoyl-L-alanine amidase sle1 OS=Staphylococcus saprophyticus subsp. saprophyticus (strain ATCC 15305 / DSM 20229 / NCIMB 8711 / NCTC 7292 / S-41) OX=342451 GN=sle1 PE=3 SV=1 |
| Q7A1T4 | 1.68e-17 | 262 | 367 | 230 | 333 | N-acetylmuramoyl-L-alanine amidase sle1 OS=Staphylococcus aureus (strain MW2) OX=196620 GN=sle1 PE=3 SV=1 |
| P0C1U7 | 1.68e-17 | 262 | 367 | 230 | 333 | N-acetylmuramoyl-L-alanine amidase sle1 OS=Staphylococcus aureus OX=1280 GN=sle1 PE=3 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as OTHER
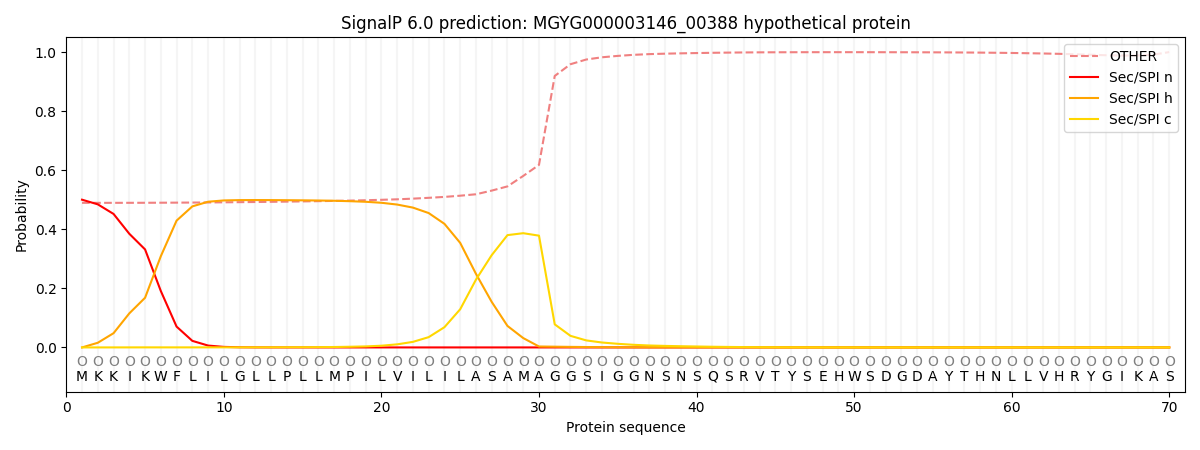
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 0.511589 | 0.469625 | 0.011847 | 0.002057 | 0.001166 | 0.003719 |