You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000003146_00453
You are here: Home > Sequence: MGYG000003146_00453
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Streptococcus oralis_S | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Firmicutes; Bacilli; Lactobacillales; Streptococcaceae; Streptococcus; Streptococcus oralis_S | |||||||||||
| CAZyme ID | MGYG000003146_00453 | |||||||||||
| CAZy Family | GH20 | |||||||||||
| CAZyme Description | Beta-N-acetylhexosaminidase | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 118382; End: 122137 Strand: + | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH20 | 161 | 511 | 1.9e-53 | 0.9317507418397626 |
| GH20 | 607 | 945 | 1.1e-50 | 0.9287833827893175 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| COG3525 | Chb | 0.0 | 152 | 939 | 6 | 732 | N-acetyl-beta-hexosaminidase [Carbohydrate transport and metabolism]. |
| cd06564 | GH20_DspB_LnbB-like | 2.23e-108 | 603 | 950 | 1 | 326 | Glycosyl hydrolase family 20 (GH20) catalytic domain of dispersin B (DspB), lacto-N-biosidase (LnbB) and related proteins. Dispersin B is a soluble beta-N-acetylglucosamidase found in bacteria that hydrolyzes the beta-1,6-linkages of PGA (poly-beta-(1,6)-N-acetylglucosamine), a major component of the extracellular polysaccharide matrix. Lacto-N-biosidase hydrolyzes lacto-N-biose (LNB) type I oligosaccharides at the nonreducing terminus to produce lacto-N-biose as part of the GNB/LNB (galacto-N-biose/lacto-N-biose I) degradation pathway. The lacto-N-biosidase from Bifidobacterium bifidum has this GH20 domain, a carbohydrate binding module 32, and a bacterial immunoglobulin-like domain 2, as well as a YSIRK signal peptide and a G5 membrane anchor at the N and C termini, respectively. The GH20 hexosaminidases are thought to act via a catalytic mechanism in which the catalytic nucleophile is not provided by solvent or the enzyme, but by the substrate itself. |
| cd06564 | GH20_DspB_LnbB-like | 8.54e-107 | 158 | 516 | 1 | 326 | Glycosyl hydrolase family 20 (GH20) catalytic domain of dispersin B (DspB), lacto-N-biosidase (LnbB) and related proteins. Dispersin B is a soluble beta-N-acetylglucosamidase found in bacteria that hydrolyzes the beta-1,6-linkages of PGA (poly-beta-(1,6)-N-acetylglucosamine), a major component of the extracellular polysaccharide matrix. Lacto-N-biosidase hydrolyzes lacto-N-biose (LNB) type I oligosaccharides at the nonreducing terminus to produce lacto-N-biose as part of the GNB/LNB (galacto-N-biose/lacto-N-biose I) degradation pathway. The lacto-N-biosidase from Bifidobacterium bifidum has this GH20 domain, a carbohydrate binding module 32, and a bacterial immunoglobulin-like domain 2, as well as a YSIRK signal peptide and a G5 membrane anchor at the N and C termini, respectively. The GH20 hexosaminidases are thought to act via a catalytic mechanism in which the catalytic nucleophile is not provided by solvent or the enzyme, but by the substrate itself. |
| COG3525 | Chb | 1.73e-37 | 597 | 980 | 6 | 379 | N-acetyl-beta-hexosaminidase [Carbohydrate transport and metabolism]. |
| cd02742 | GH20_hexosaminidase | 4.85e-28 | 607 | 885 | 4 | 249 | Beta-N-acetylhexosaminidases of glycosyl hydrolase family 20 (GH20) catalyze the removal of beta-1,4-linked N-acetyl-D-hexosamine residues from the non-reducing ends of N-acetyl-beta-D-hexosaminides including N-acetylglucosides and N-acetylgalactosides. These enzymes are broadly distributed in microorganisms, plants and animals, and play roles in various key physiological and pathological processes. These processes include cell structural integrity, energy storage, cellular signaling, fertilization, pathogen defense, viral penetration, the development of carcinomas, inflammatory events and lysosomal storage disorders. The GH20 enzymes include the eukaryotic beta-N-acetylhexosaminidases A and B, the bacterial chitobiases, dispersin B, and lacto-N-biosidase. The GH20 hexosaminidases are thought to act via a catalytic mechanism in which the catalytic nucleophile is not provided by the solvent or the enzyme, but by the substrate itself. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| QLL96593.1 | 0.0 | 1 | 1251 | 1 | 1251 |
| ATF57646.1 | 0.0 | 1 | 1251 | 1 | 1251 |
| ANR74964.1 | 0.0 | 1 | 1251 | 1 | 1251 |
| QLL97743.1 | 0.0 | 1 | 1251 | 1 | 1251 |
| VEF78162.1 | 0.0 | 1 | 1251 | 1 | 1251 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 3RPM_A | 1.72e-292 | 148 | 614 | 1 | 467 | Crystalstructure of the first GH20 domain of a novel Beta-N-acetyl-hexosaminidase StrH from Streptococcus pneumoniae R6 [Streptococcus pneumoniae R6],3RPM_B Crystal structure of the first GH20 domain of a novel Beta-N-acetyl-hexosaminidase StrH from Streptococcus pneumoniae R6 [Streptococcus pneumoniae R6] |
| 4AZI_A | 1.07e-283 | 598 | 1036 | 4 | 442 | Differentialinhibition of the tandem GH20 catalytic modules in the pneumococcal exo-beta-D-N-acetylglucosaminidase, StrH [Streptococcus pneumoniae TIGR4],4AZI_B Differential inhibition of the tandem GH20 catalytic modules in the pneumococcal exo-beta-D-N-acetylglucosaminidase, StrH [Streptococcus pneumoniae TIGR4] |
| 2YL5_A | 1.22e-282 | 598 | 1036 | 4 | 442 | Inhibitionof the pneumococcal virulence factor StrH and molecular insights into N-glycan recognition and hydrolysis [Streptococcus pneumoniae TIGR4],2YL5_B Inhibition of the pneumococcal virulence factor StrH and molecular insights into N-glycan recognition and hydrolysis [Streptococcus pneumoniae TIGR4],2YL5_C Inhibition of the pneumococcal virulence factor StrH and molecular insights into N-glycan recognition and hydrolysis [Streptococcus pneumoniae TIGR4],2YL5_D Inhibition of the pneumococcal virulence factor StrH and molecular insights into N-glycan recognition and hydrolysis [Streptococcus pneumoniae TIGR4],4AZC_B Differential inhibition of the tandem GH20 catalytic modules in the pneumococcal exo-beta-D-N-acetylglucosaminidase, StrH [Streptococcus pneumoniae TIGR4],4AZC_C Differential inhibition of the tandem GH20 catalytic modules in the pneumococcal exo-beta-D-N-acetylglucosaminidase, StrH [Streptococcus pneumoniae TIGR4],4AZC_D Differential inhibition of the tandem GH20 catalytic modules in the pneumococcal exo-beta-D-N-acetylglucosaminidase, StrH [Streptococcus pneumoniae TIGR4] |
| 4AZG_A | 1.26e-282 | 598 | 1036 | 5 | 443 | Differentialinhibition of the tandem GH20 catalytic modules in the pneumococcal exo-beta-D-N-acetylglucosaminidase, StrH [Streptococcus pneumoniae TIGR4],4AZG_B Differential inhibition of the tandem GH20 catalytic modules in the pneumococcal exo-beta-D-N-acetylglucosaminidase, StrH [Streptococcus pneumoniae TIGR4],4AZH_A Differential inhibition of the tandem GH20 catalytic modules in the pneumococcal exo-beta-D-N-acetylglucosaminidase, StrH [Streptococcus pneumoniae TIGR4],4AZH_B Differential inhibition of the tandem GH20 catalytic modules in the pneumococcal exo-beta-D-N-acetylglucosaminidase, StrH [Streptococcus pneumoniae TIGR4],4AZH_C Differential inhibition of the tandem GH20 catalytic modules in the pneumococcal exo-beta-D-N-acetylglucosaminidase, StrH [Streptococcus pneumoniae TIGR4],4AZH_D Differential inhibition of the tandem GH20 catalytic modules in the pneumococcal exo-beta-D-N-acetylglucosaminidase, StrH [Streptococcus pneumoniae TIGR4] |
| 2YL6_A | 2.55e-282 | 153 | 586 | 1 | 434 | Inhibitionof the pneumococcal virulence factor StrH and molecular insights into N-glycan recognition and hydrolysis [Streptococcus pneumoniae TIGR4],4AZ5_A Differential inhibition of the tandem GH20 catalytic modules in the pneumococcal exo-beta-D-N-acetylglucosaminidase, StrH [Streptococcus pneumoniae TIGR4] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| P49610 | 0.0 | 1 | 1251 | 1 | 1312 | Beta-N-acetylhexosaminidase OS=Streptococcus pneumoniae serotype 4 (strain ATCC BAA-334 / TIGR4) OX=170187 GN=strH PE=1 SV=2 |
| B2UPR7 | 7.54e-09 | 164 | 491 | 239 | 538 | Beta-hexosaminidase Amuc_2136 OS=Akkermansia muciniphila (strain ATCC BAA-835 / DSM 22959 / JCM 33894 / BCRC 81048 / CCUG 64013 / CIP 107961 / Muc) OX=349741 GN=Amuc_2136 PE=1 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as SP
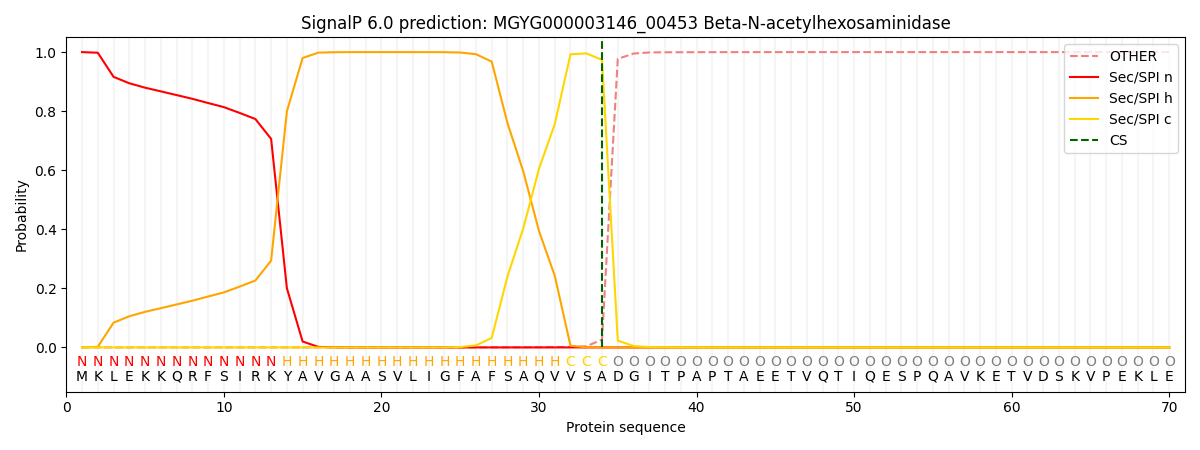
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 0.000317 | 0.998976 | 0.000198 | 0.000186 | 0.000174 | 0.000145 |
