You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000003199_00026
You are here: Home > Sequence: MGYG000003199_00026
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | HGM05190 sp900759815 | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Bacteroidota; Bacteroidia; Bacteroidales; Muribaculaceae; HGM05190; HGM05190 sp900759815 | |||||||||||
| CAZyme ID | MGYG000003199_00026 | |||||||||||
| CAZy Family | GH28 | |||||||||||
| CAZyme Description | hypothetical protein | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 8101; End: 9453 Strand: - | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH28 | 79 | 422 | 5.7e-59 | 0.916923076923077 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| COG5434 | Pgu1 | 1.10e-48 | 28 | 354 | 59 | 395 | Polygalacturonase [Carbohydrate transport and metabolism]. |
| pfam00295 | Glyco_hydro_28 | 1.06e-19 | 85 | 434 | 8 | 319 | Glycosyl hydrolases family 28. Glycosyl hydrolase family 28 includes polygalacturonase EC:3.2.1.15 as well as rhamnogalacturonase A(RGase A), EC:3.2.1.-. These enzymes are important in cell wall metabolism. |
| PLN03003 | PLN03003 | 1.03e-09 | 152 | 335 | 117 | 269 | Probable polygalacturonase At3g15720 |
| PLN03010 | PLN03010 | 3.04e-09 | 52 | 353 | 47 | 306 | polygalacturonase |
| pfam13229 | Beta_helix | 3.85e-06 | 194 | 329 | 3 | 138 | Right handed beta helix region. This region contains a parallel beta helix region that shares some similarity with Pectate lyases. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| QCP73449.1 | 4.16e-195 | 51 | 442 | 340 | 731 |
| QCD39802.1 | 4.16e-195 | 51 | 442 | 340 | 731 |
| QCD43023.1 | 3.18e-192 | 43 | 442 | 333 | 731 |
| QNR82966.1 | 2.10e-168 | 46 | 441 | 16 | 410 |
| ARS40669.1 | 1.71e-167 | 52 | 441 | 22 | 410 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 5OLP_A | 2.37e-35 | 48 | 417 | 41 | 424 | Galacturonidase[Bacteroides thetaiotaomicron VPI-5482],5OLP_B Galacturonidase [Bacteroides thetaiotaomicron VPI-5482] |
| 3JUR_A | 1.45e-30 | 48 | 441 | 24 | 433 | Thecrystal structure of a hyperthermoactive Exopolygalacturonase from Thermotoga maritima [Thermotoga maritima],3JUR_B The crystal structure of a hyperthermoactive Exopolygalacturonase from Thermotoga maritima [Thermotoga maritima],3JUR_C The crystal structure of a hyperthermoactive Exopolygalacturonase from Thermotoga maritima [Thermotoga maritima],3JUR_D The crystal structure of a hyperthermoactive Exopolygalacturonase from Thermotoga maritima [Thermotoga maritima] |
| 4MXN_A | 7.71e-16 | 50 | 263 | 20 | 215 | Crystalstructure of a putative glycosyl hydrolase (PARMER_00599) from Parabacteroides merdae ATCC 43184 at 1.95 A resolution [Parabacteroides merdae ATCC 43184],4MXN_B Crystal structure of a putative glycosyl hydrolase (PARMER_00599) from Parabacteroides merdae ATCC 43184 at 1.95 A resolution [Parabacteroides merdae ATCC 43184],4MXN_C Crystal structure of a putative glycosyl hydrolase (PARMER_00599) from Parabacteroides merdae ATCC 43184 at 1.95 A resolution [Parabacteroides merdae ATCC 43184],4MXN_D Crystal structure of a putative glycosyl hydrolase (PARMER_00599) from Parabacteroides merdae ATCC 43184 at 1.95 A resolution [Parabacteroides merdae ATCC 43184] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| A7PZL3 | 1.41e-21 | 54 | 424 | 65 | 430 | Probable polygalacturonase OS=Vitis vinifera OX=29760 GN=GSVIVT00026920001 PE=1 SV=1 |
| P15922 | 3.38e-11 | 54 | 445 | 154 | 601 | Exo-poly-alpha-D-galacturonosidase OS=Dickeya chrysanthemi OX=556 GN=pehX PE=1 SV=1 |
| P35338 | 2.20e-10 | 35 | 363 | 26 | 325 | Exopolygalacturonase OS=Zea mays OX=4577 GN=PG9 PE=2 SV=1 |
| Q9LW07 | 4.45e-10 | 152 | 353 | 117 | 287 | Probable polygalacturonase At3g15720 OS=Arabidopsis thaliana OX=3702 GN=At3g15720 PE=3 SV=1 |
| Q949Z1 | 1.44e-09 | 54 | 352 | 82 | 363 | Polygalacturonase At1g48100 OS=Arabidopsis thaliana OX=3702 GN=At1g48100 PE=2 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as SP
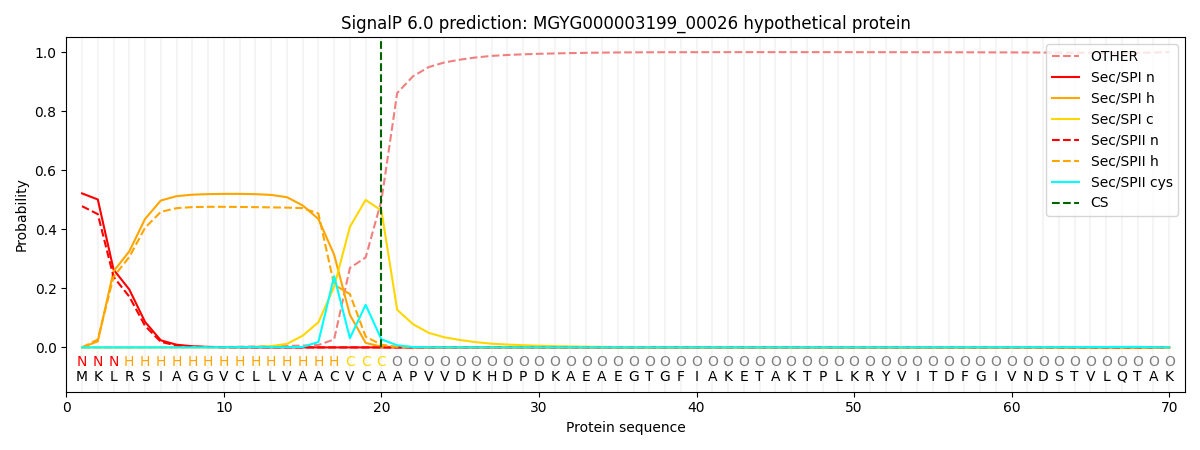
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 0.001062 | 0.510471 | 0.487659 | 0.000266 | 0.000288 | 0.000237 |
