You are browsing environment: HUMAN GUT
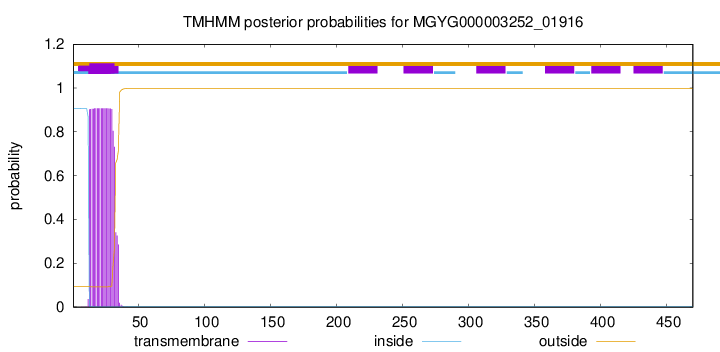
MGYG000003252_01916
Basic Information
help
Species
Bacteroides sp900761785
Lineage
Bacteria; Bacteroidota; Bacteroidia; Bacteroidales; Bacteroidaceae; Bacteroides; Bacteroides sp900761785
CAZyme ID
MGYG000003252_01916
CAZy Family
GH28
CAZyme Description
hypothetical protein
CAZyme Property
Genome Property
Genome Assembly ID
Genome Size
Genome Type
Country
Continent
MGYG000003252
8296673
MAG
United States
North America
Gene Location
Start: 93440;
End: 94852
Strand: -
No EC number prediction in MGYG000003252_01916.
Family
Start
End
Evalue
family coverage
GH28
177
455
3.7e-40
0.803076923076923
Cdd ID
Domain
E-Value
qStart
qEnd
sStart
sEnd
Domain Description
COG5434
Pgu1
5.59e-11
218
352
239
371
Polygalacturonase [Carbohydrate transport and metabolism].
more
pfam00295
Glyco_hydro_28
4.71e-06
197
451
88
307
Glycosyl hydrolases family 28. Glycosyl hydrolase family 28 includes polygalacturonase EC:3.2.1.15 as well as rhamnogalacturonase A(RGase A), EC:3.2.1.-. These enzymes are important in cell wall metabolism.
more
PLN02793
PLN02793
0.003
219
383
229
396
Probable polygalacturonase
more
This protein is predicted as SP
Other
SP_Sec_SPI
LIPO_Sec_SPII
TAT_Tat_SPI
TATLIP_Sec_SPII
PILIN_Sec_SPIII
0.441974
0.543982
0.009399
0.002670
0.000898
0.001047