You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000003279_00303
You are here: Home > Sequence: MGYG000003279_00303
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Alistipes sp900541585 | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Bacteroidota; Bacteroidia; Bacteroidales; Rikenellaceae; Alistipes; Alistipes sp900541585 | |||||||||||
| CAZyme ID | MGYG000003279_00303 | |||||||||||
| CAZy Family | GT32 | |||||||||||
| CAZyme Description | hypothetical protein | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 46892; End: 47746 Strand: - | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GT32 | 26 | 100 | 5.7e-21 | 0.9222222222222223 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| COG3774 | OCH1 | 1.54e-16 | 2 | 103 | 77 | 181 | Mannosyltransferase OCH1 or related enzyme [Cell wall/membrane/envelope biogenesis]. |
| pfam05704 | Caps_synth | 1.16e-13 | 2 | 106 | 41 | 151 | Capsular polysaccharide synthesis protein. This family consists of several capsular polysaccharide proteins. Capsular polysaccharide (CPS) is a major virulence factor in Streptococcus pneumoniae. |
| pfam04488 | Gly_transf_sug | 1.90e-11 | 22 | 105 | 1 | 92 | Glycosyltransferase sugar-binding region containing DXD motif. The DXD motif is a short conserved motif found in many families of glycosyltransferases, which add a range of different sugars to other sugars, phosphates and proteins. DXD-containing glycosyltransferases all use nucleoside diphosphate sugars as donors and require divalent cations, usually manganese. The DXD motif is expected to play a carbohydrate binding role in sugar-nucleoside diphosphate and manganese dependent glycosyltransferases. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| QUT47110.1 | 7.14e-71 | 7 | 237 | 3 | 233 |
| CBK68805.1 | 1.39e-69 | 7 | 235 | 2 | 231 |
| QUR43722.1 | 1.39e-69 | 7 | 235 | 2 | 231 |
| QRM97945.1 | 1.39e-69 | 7 | 235 | 2 | 231 |
| BBA29457.1 | 2.32e-67 | 7 | 235 | 2 | 231 |
Swiss-Prot Hits help
SignalP and Lipop Annotations help
This protein is predicted as OTHER
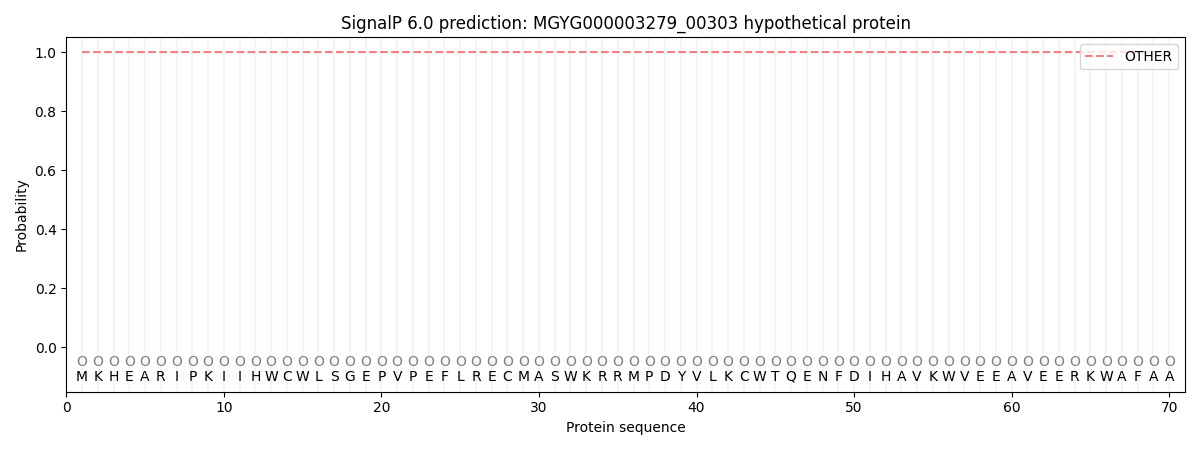
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 1.000059 | 0.000000 | 0.000000 | 0.000000 | 0.000000 | 0.000000 |
