You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000003346_02631
You are here: Home > Sequence: MGYG000003346_02631
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
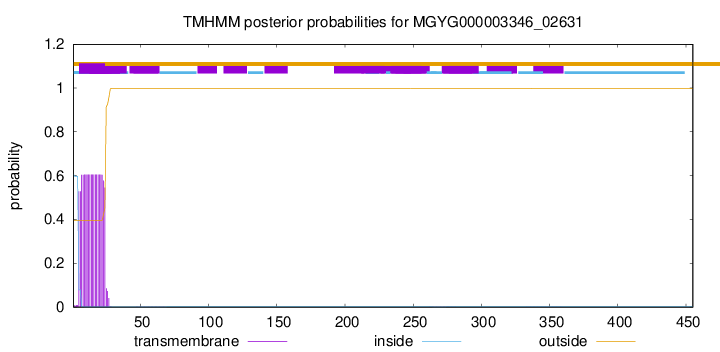
TMHMM annotations
Basic Information help
| Species | Prevotella sp900554545 | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Bacteroidota; Bacteroidia; Bacteroidales; Bacteroidaceae; Prevotella; Prevotella sp900554545 | |||||||||||
| CAZyme ID | MGYG000003346_02631 | |||||||||||
| CAZy Family | CE3 | |||||||||||
| CAZyme Description | hypothetical protein | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 16892; End: 18259 Strand: + | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| CE3 | 32 | 221 | 2e-42 | 0.9896907216494846 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| cd01833 | XynB_like | 1.18e-25 | 31 | 221 | 1 | 157 | SGNH_hydrolase subfamily, similar to Ruminococcus flavefaciens XynB. Most likely a secreted hydrolase with xylanase activity. SGNH hydrolases are a diverse family of lipases and esterases. The tertiary fold of the enzyme is substantially different from that of the alpha/beta hydrolase family and unique among all known hydrolases; its active site closely resembles the Ser-His-Asp(Glu) triad found in other serine hydrolases. |
| pfam13472 | Lipase_GDSL_2 | 7.11e-18 | 36 | 209 | 2 | 173 | GDSL-like Lipase/Acylhydrolase family. This family of presumed lipases and related enzymes are similar to pfam00657. |
| cd00229 | SGNH_hydrolase | 8.82e-16 | 36 | 220 | 4 | 187 | SGNH_hydrolase, or GDSL_hydrolase, is a diverse family of lipases and esterases. The tertiary fold of the enzyme is substantially different from that of the alpha/beta hydrolase family and unique among all known hydrolases; its active site closely resembles the typical Ser-His-Asp(Glu) triad from other serine hydrolases, but may lack the carboxlic acid. |
| cd01828 | sialate_O-acetylesterase_like2 | 1.55e-14 | 36 | 225 | 5 | 168 | sialate_O-acetylesterase_like subfamily of the SGNH-hydrolases, a diverse family of lipases and esterases. The tertiary fold of the enzyme is substantially different from that of the alpha/beta hydrolase family and unique among all known hydrolases; its active site closely resembles the Ser-His-Asp(Glu) triad found in other serine hydrolases. |
| COG2755 | TesA | 5.24e-11 | 32 | 225 | 10 | 212 | Lysophospholipase L1 or related esterase [Amino acid transport and metabolism]. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| BAX78799.1 | 5.52e-116 | 25 | 445 | 32 | 488 |
| ATC65549.1 | 1.22e-112 | 6 | 430 | 15 | 474 |
| QOV88633.1 | 2.62e-62 | 14 | 227 | 13 | 228 |
| QNN20991.1 | 9.05e-51 | 23 | 221 | 22 | 222 |
| QRK14019.1 | 7.37e-27 | 34 | 225 | 1 | 199 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 5AO9_A | 1.93e-23 | 236 | 446 | 18 | 273 | Thestructure of a novel thermophilic esterase from the Planctomycetes species, Thermogutta terrifontis, Est2-native [Thermogutta terrifontis],5AOA_A The structure of a novel thermophilic esterase from the Planctomycetes species, Thermogutta terrifontis, Est2-Propionate bound [Thermogutta terrifontis],5AOB_A The structure of a novel thermophilic esterase from the Planctomycetes species, Thermogutta terrifontis, Est2-butyrate bound [Thermogutta terrifontis],5AOC_A The structure of a novel thermophilic esterase from the Planctomycetes species, Thermogutta terrifontis, Est2-valerate bound [Thermogutta terrifontis] |
| 7BFN_A | 1.97e-23 | 236 | 446 | 19 | 274 | ChainA, Esterase [Thermogutta terrifontis] |
| 7BFO_A | 9.38e-23 | 236 | 446 | 19 | 274 | ChainA, Esterase [Thermogutta terrifontis],7BFR_A Chain A, Esterase [Thermogutta terrifontis],7BFT_A Chain A, Esterase [Thermogutta terrifontis],7BFU_A Chain A, Esterase [Thermogutta terrifontis],7BFV_A Chain A, Esterase [Thermogutta terrifontis] |
| 4YPV_A | 4.16e-09 | 214 | 369 | 67 | 229 | High-resolutionstructure of a metagenome-derived esterase Est8 [Parvibaculum] |
| 2YH2_A | 1.95e-08 | 245 | 368 | 59 | 189 | Pyrobaculumcalidifontis esterase monoclinic form [Pyrobaculum calidifontis],2YH2_B Pyrobaculum calidifontis esterase monoclinic form [Pyrobaculum calidifontis],2YH2_C Pyrobaculum calidifontis esterase monoclinic form [Pyrobaculum calidifontis],2YH2_D Pyrobaculum calidifontis esterase monoclinic form [Pyrobaculum calidifontis],3ZWQ_A Hyperthermophilic Esterase From The Archeon Pyrobaculum Calidifontis [Pyrobaculum calidifontis JCM 11548],3ZWQ_B Hyperthermophilic Esterase From The Archeon Pyrobaculum Calidifontis [Pyrobaculum calidifontis JCM 11548] |
Swiss-Prot Hits help
SignalP and Lipop Annotations help
This protein is predicted as SP
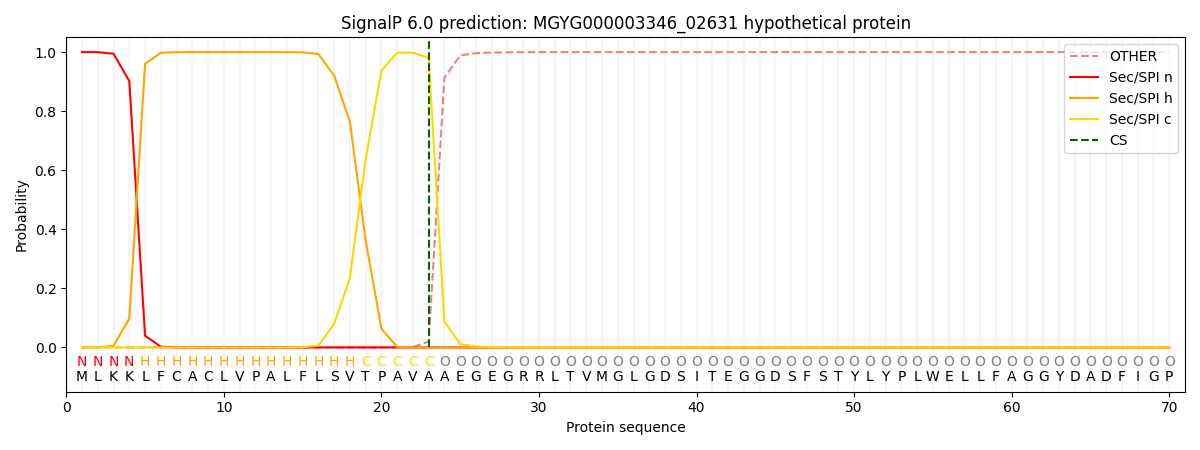
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 0.000241 | 0.999006 | 0.000214 | 0.000165 | 0.000165 | 0.000154 |