You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000003362_01157
You are here: Home > Sequence: MGYG000003362_01157
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Dysgonomonas sp900079735 | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Bacteroidota; Bacteroidia; Bacteroidales; Dysgonomonadaceae; Dysgonomonas; Dysgonomonas sp900079735 | |||||||||||
| CAZyme ID | MGYG000003362_01157 | |||||||||||
| CAZy Family | GT80 | |||||||||||
| CAZyme Description | hypothetical protein | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 109461; End: 111170 Strand: - | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GT80 | 308 | 549 | 1.2e-24 | 0.6490765171503958 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| cd14948 | BACON | 2.50e-09 | 23 | 102 | 2 | 82 | Bacteroidetes-Associated Carbohydrate-binding (putative) Often N-terminal (BACON) domain. The BACON domain is found in diverse domain architectures and accociated with a wide variety of domains, including carbohydrate-active enzymes and proteases. It was named for its suggested function of carbohydrate binding; the latter was inferred from domain architectures, sequence conservation, and phyletic distribution. However, recent experimental data suggest that its primary function in Bacteroides ovatus endo-xyloglucanase BoGH5A is to distance the catalytic module from the cell surface and confer additional mobility to the catalytic domain for attack of the polysaccharide. No evidence for a direct role in carbohydrate binding could be found in that case. The large majority of BACON domains are found in Bacteroidetes. |
| cd14948 | BACON | 2.18e-07 | 108 | 192 | 1 | 82 | Bacteroidetes-Associated Carbohydrate-binding (putative) Often N-terminal (BACON) domain. The BACON domain is found in diverse domain architectures and accociated with a wide variety of domains, including carbohydrate-active enzymes and proteases. It was named for its suggested function of carbohydrate binding; the latter was inferred from domain architectures, sequence conservation, and phyletic distribution. However, recent experimental data suggest that its primary function in Bacteroides ovatus endo-xyloglucanase BoGH5A is to distance the catalytic module from the cell surface and confer additional mobility to the catalytic domain for attack of the polysaccharide. No evidence for a direct role in carbohydrate binding could be found in that case. The large majority of BACON domains are found in Bacteroidetes. |
| pfam19190 | BACON_2 | 9.24e-05 | 118 | 185 | 9 | 82 | Viral BACON domain. This family represents a distinct class of BACON domains found in crAss-like phages, the most common viral family in the human gut, in which they are found in tail fiber genes. This suggests they may play a role in phage-host interactions. |
| pfam13004 | BACON | 0.003 | 136 | 192 | 2 | 60 | Putative binding domain, N-terminal. The BACON (Bacteroidetes-Associated Carbohydrate-binding Often N-terminal) domain is an all-beta domain found in diverse architectures, principally in combination with carbohydrate-active enzymes and proteases. These architectures suggest a carbohydrate-binding function which is also supported by the nature of BACON's few conserved amino-acids. The phyletic distribution of BACON and other data tentatively suggest that it may frequently function to bind mucin. Further work with the characterized structure of a member of glycoside hydrolase family 5 enzyme, Structure 3ZMR, has found no evidence for carbohydrate-binding for this domain. |
| pfam13004 | BACON | 0.006 | 48 | 102 | 2 | 60 | Putative binding domain, N-terminal. The BACON (Bacteroidetes-Associated Carbohydrate-binding Often N-terminal) domain is an all-beta domain found in diverse architectures, principally in combination with carbohydrate-active enzymes and proteases. These architectures suggest a carbohydrate-binding function which is also supported by the nature of BACON's few conserved amino-acids. The phyletic distribution of BACON and other data tentatively suggest that it may frequently function to bind mucin. Further work with the characterized structure of a member of glycoside hydrolase family 5 enzyme, Structure 3ZMR, has found no evidence for carbohydrate-binding for this domain. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| BBL01105.1 | 1.56e-130 | 106 | 567 | 27 | 497 |
| BBL09010.1 | 5.46e-45 | 385 | 567 | 2 | 188 |
| BBL11802.1 | 5.46e-45 | 385 | 567 | 2 | 188 |
| BAA25316.1 | 4.70e-12 | 311 | 562 | 224 | 480 |
| AWK81336.1 | 1.03e-10 | 311 | 562 | 224 | 480 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 4R9V_A | 8.69e-14 | 311 | 562 | 133 | 389 | Crystalstructure of sialyltransferase from photobacterium damselae, residues 113-497 corresponding to the gt-b domain [Photobacterium damselae] |
| 4R83_A | 1.27e-13 | 311 | 562 | 230 | 486 | Crystalstructure of Sialyltransferase from Photobacterium damsela [Photobacterium damselae],4R83_B Crystal structure of Sialyltransferase from Photobacterium damsela [Photobacterium damselae],4R83_C Crystal structure of Sialyltransferase from Photobacterium damsela [Photobacterium damselae],4R83_D Crystal structure of Sialyltransferase from Photobacterium damsela [Photobacterium damselae],4R84_A Crystal structure of Sialyltransferase from Photobacterium damsela with CMP-3F(a)Neu5Ac bound [Photobacterium damselae],4R84_B Crystal structure of Sialyltransferase from Photobacterium damsela with CMP-3F(a)Neu5Ac bound [Photobacterium damselae],4R84_C Crystal structure of Sialyltransferase from Photobacterium damsela with CMP-3F(a)Neu5Ac bound [Photobacterium damselae],4R84_D Crystal structure of Sialyltransferase from Photobacterium damsela with CMP-3F(a)Neu5Ac bound [Photobacterium damselae] |
| 2Z4T_A | 8.83e-12 | 311 | 550 | 227 | 472 | CrystalStructure of Vibrionaceae Photobacterium sp. JT-ISH-224 2,6-sialyltransferase in a Ternary Complex with Donor Product CMP and Accepter Substrate Lactose [Photobacterium sp. JT-ISH-224] |
Swiss-Prot Hits help
SignalP and Lipop Annotations help
This protein is predicted as OTHER
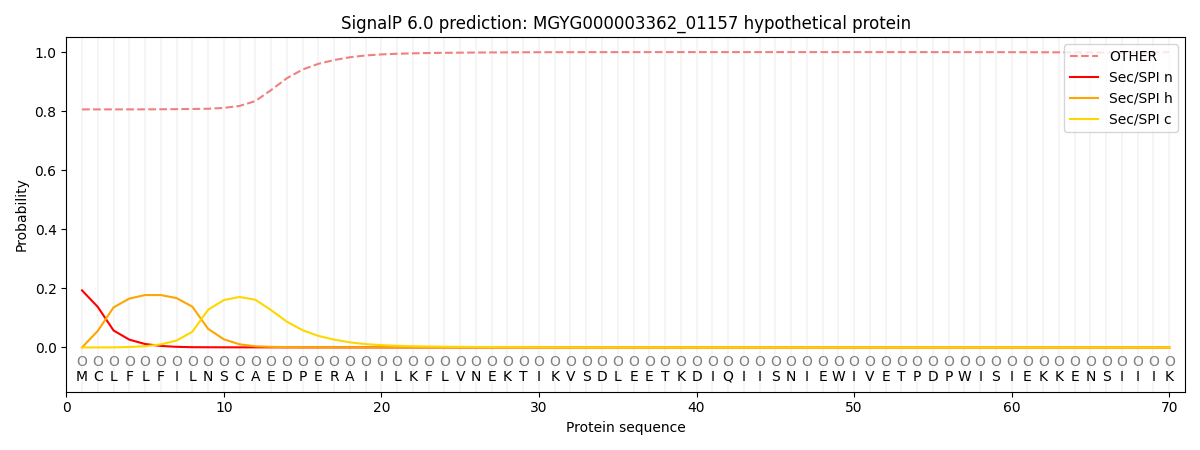
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 0.817974 | 0.179604 | 0.001353 | 0.000408 | 0.000216 | 0.000457 |
