You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000003377_04592
You are here: Home > Sequence: MGYG000003377_04592
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Phytobacter sp002377245 | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Proteobacteria; Gammaproteobacteria; Enterobacterales; Enterobacteriaceae; Phytobacter; Phytobacter sp002377245 | |||||||||||
| CAZyme ID | MGYG000003377_04592 | |||||||||||
| CAZy Family | PL5 | |||||||||||
| CAZyme Description | Alginate lyase | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 125820; End: 126902 Strand: - | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| PL5 | 24 | 336 | 6.4e-138 | 0.9810126582278481 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| PRK00325 | algL | 7.06e-136 | 16 | 357 | 17 | 355 | polysaccharide lyase. |
| cd00244 | AlgLyase | 1.54e-132 | 19 | 357 | 2 | 339 | Alginate Lyase A1-III; enzymatically depolymerizes alginate, a complex copolymer of beta-D-mannuronate and alpha-L-guluronate, by cleaving the beta-(1,4) glycosidic bond. |
| pfam05426 | Alginate_lyase | 6.55e-40 | 59 | 299 | 2 | 274 | Alginate lyase. This family contains several bacterial alginate lyase proteins. Alginate is a family of 1-4-linked copolymers of beta -D-mannuronic acid (M) and alpha -L-guluronic acid (G). It is produced by brown algae and by some bacteria belonging to the genera Azotobacter and Pseudomonas. Alginate lyases catalyze the depolymerization of alginates by beta -elimination, generating a molecule containing 4-deoxy-L-erythro-hex-4-enepyranosyluronate at the nonreducing end. This family adopts an all alpha fold. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| QJF17808.1 | 1.35e-160 | 1 | 358 | 1 | 356 |
| AUV08955.1 | 3.87e-160 | 1 | 358 | 1 | 356 |
| AUU91003.1 | 3.87e-160 | 1 | 358 | 1 | 356 |
| QIH63616.1 | 7.78e-160 | 1 | 358 | 1 | 356 |
| BBE77835.1 | 9.31e-159 | 12 | 358 | 2 | 347 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 4OZV_A | 8.02e-114 | 24 | 358 | 3 | 335 | CrystalStructure of the periplasmic alginate lyase AlgL [Pseudomonas aeruginosa PAO1] |
| 4OZW_A | 2.62e-112 | 24 | 358 | 3 | 335 | CrystalStructure of the periplasmic alginate lyase AlgL H202A mutant [Pseudomonas aeruginosa PAO1] |
| 7FHZ_A | 7.14e-38 | 67 | 356 | 20 | 299 | ChainA, Polysaccharide lyase [Stenotrophomonas maltophilia K279a] |
| 7FHU_A | 8.37e-38 | 67 | 356 | 20 | 299 | ChainA, Polysaccharide lyase [Stenotrophomonas maltophilia K279a],7FHU_B Chain B, Polysaccharide lyase [Stenotrophomonas maltophilia K279a],7FHV_A Chain A, Polysaccharide lyase [Stenotrophomonas maltophilia K279a],7FHV_B Chain B, Polysaccharide lyase [Stenotrophomonas maltophilia K279a],7FHW_A Chain A, Polysaccharide lyase [Stenotrophomonas maltophilia K279a],7FHW_B Chain B, Polysaccharide lyase [Stenotrophomonas maltophilia K279a],7FHX_A Chain A, Polysaccharide lyase [Stenotrophomonas maltophilia K279a],7FHX_B Chain B, Polysaccharide lyase [Stenotrophomonas maltophilia K279a],7FHY_A Chain A, Polysaccharide lyase [Stenotrophomonas maltophilia K279a],7FHY_B Chain B, Polysaccharide lyase [Stenotrophomonas maltophilia K279a],7FI0_A Chain A, Polysaccharide lyase [Stenotrophomonas maltophilia K279a],7FI1_A Chain A, Polysaccharide lyase [Stenotrophomonas maltophilia K279a] |
| 7FI2_A | 2.35e-36 | 67 | 356 | 20 | 299 | ChainA, Polysaccharide lyase [Stenotrophomonas maltophilia K279a] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| Q4KHY5 | 7.31e-118 | 13 | 357 | 19 | 363 | Alginate lyase OS=Pseudomonas fluorescens (strain ATCC BAA-477 / NRRL B-23932 / Pf-5) OX=220664 GN=algL PE=3 SV=1 |
| Q9L7P2 | 2.12e-116 | 20 | 357 | 26 | 363 | Alginate lyase OS=Pseudomonas syringae pv. syringae OX=321 GN=algL PE=1 SV=1 |
| A6V1P7 | 8.42e-116 | 18 | 358 | 25 | 362 | Alginate lyase OS=Pseudomonas aeruginosa (strain PA7) OX=381754 GN=algL PE=3 SV=1 |
| Q4ZXL0 | 8.53e-116 | 20 | 357 | 26 | 363 | Alginate lyase OS=Pseudomonas syringae pv. syringae (strain B728a) OX=205918 GN=algL PE=3 SV=1 |
| B1J479 | 2.40e-115 | 10 | 357 | 12 | 359 | Alginate lyase OS=Pseudomonas putida (strain W619) OX=390235 GN=algL PE=3 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as SP
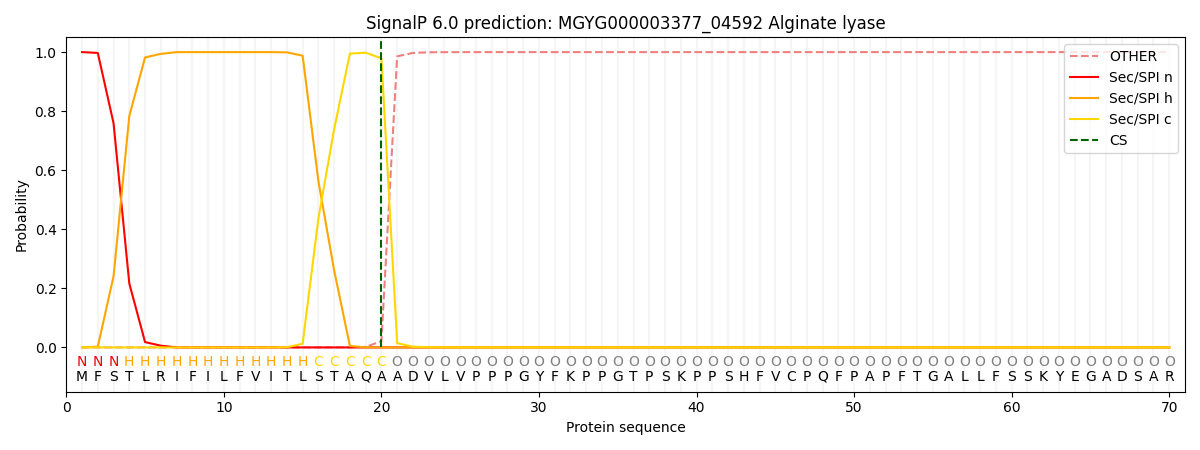
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 0.000217 | 0.999228 | 0.000148 | 0.000142 | 0.000131 | 0.000128 |
