You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000003403_00060
You are here: Home > Sequence: MGYG000003403_00060
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Stenotrophomonas maltophilia_AM | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Proteobacteria; Gammaproteobacteria; Xanthomonadales; Xanthomonadaceae; Stenotrophomonas; Stenotrophomonas maltophilia_AM | |||||||||||
| CAZyme ID | MGYG000003403_00060 | |||||||||||
| CAZy Family | GH3 | |||||||||||
| CAZyme Description | hypothetical protein | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 60087; End: 62663 Strand: + | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH3 | 145 | 365 | 1e-67 | 0.9583333333333334 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| COG1472 | BglX | 8.83e-97 | 76 | 502 | 1 | 392 | Periplasmic beta-glucosidase and related glycosidases [Carbohydrate transport and metabolism]. |
| pfam00933 | Glyco_hydro_3 | 1.12e-79 | 77 | 401 | 1 | 316 | Glycosyl hydrolase family 3 N terminal domain. |
| PRK15098 | PRK15098 | 3.41e-79 | 50 | 656 | 20 | 648 | beta-glucosidase BglX. |
| pfam01915 | Glyco_hydro_3_C | 2.68e-38 | 442 | 656 | 1 | 216 | Glycosyl hydrolase family 3 C-terminal domain. This domain is involved in catalysis and may be involved in binding beta-glucan. This domain is found associated with pfam00933. |
| PLN03080 | PLN03080 | 1.70e-34 | 68 | 656 | 52 | 633 | Probable beta-xylosidase; Provisional |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| VEE51477.1 | 0.0 | 1 | 858 | 1 | 858 |
| QPX92218.1 | 0.0 | 1 | 858 | 1 | 858 |
| QGL93037.1 | 0.0 | 1 | 858 | 1 | 858 |
| AYZ70231.1 | 0.0 | 1 | 858 | 1 | 858 |
| QJP19971.1 | 0.0 | 1 | 858 | 1 | 858 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 3USZ_A | 5.30e-253 | 51 | 845 | 8 | 799 | Crystalstructure of truncated exo-1,3/1,4-beta-glucanase (EXOP) from Pseudoalteromonas sp. BB1 [Pseudoalteromonas sp. BB1],3UT0_A Crystal structure of exo-1,3/1,4-beta-glucanase (EXOP) from Pseudoalteromonas sp. BB1 [Pseudoalteromonas sp. BB1],3UT0_B Crystal structure of exo-1,3/1,4-beta-glucanase (EXOP) from Pseudoalteromonas sp. BB1 [Pseudoalteromonas sp. BB1],3UT0_C Crystal structure of exo-1,3/1,4-beta-glucanase (EXOP) from Pseudoalteromonas sp. BB1 [Pseudoalteromonas sp. BB1],3UT0_D Crystal structure of exo-1,3/1,4-beta-glucanase (EXOP) from Pseudoalteromonas sp. BB1 [Pseudoalteromonas sp. BB1] |
| 3RRX_A | 4.26e-252 | 51 | 845 | 8 | 799 | CrystalStructure of Q683A mutant of Exo-1,3/1,4-beta-glucanase (ExoP) from Pseudoalteromonas sp. BB1 [Pseudoalteromonas sp. BB1] |
| 5M6G_A | 9.64e-138 | 47 | 657 | 25 | 615 | Crystalstructure Glucan 1,4-beta-glucosidase from Saccharopolyspora erythraea [Saccharopolyspora erythraea D] |
| 1LQ2_A | 3.31e-134 | 65 | 655 | 12 | 598 | Crystalstructure of barley beta-D-glucan glucohydrolase isoenzyme Exo1 in complex with gluco-phenylimidazole [Hordeum vulgare],1X38_A crystal structure of barley beta-D-glucan glucohydrolase isoenzyme exo1 in complex with gluco-phenylimidazole [Hordeum vulgare],1X39_A Crystal structure of barley beta-D-glucan glucohydrolase isoenzyme exo1 in complex with gluco-phenylimidazole [Hordeum vulgare] |
| 1EX1_A | 3.63e-134 | 65 | 655 | 12 | 598 | Beta-d-glucanExohydrolase From Barley [Hordeum vulgare],1IEQ_A Crystal Structure Of Barley Beta-D-Glucan Glucohydrolase Isoenzyme Exo1 [Hordeum vulgare],1IEV_A Crystal Structure Of Barley Beta-d-glucan Glucohydrolase Isoenzyme Exo1 In Complex With Cyclohexitol [Hordeum vulgare],1IEW_A Crystal structure of barley beta-D-glucan glucohydrolase isoenzyme Exo1 in complex with 2-deoxy-2-fluoro-alpha-D-glucoside [Hordeum vulgare],1IEX_A Crystal structure of barley beta-D-glucan glucohydrolase isoenzyme Exo1 in complex with 4I,4III,4V-S-trithiocellohexaose [Hordeum vulgare],1J8V_A Crystal structure of barley beta-D-glucan glucohydrolase isoenzyme Exo1 in complex with 4'-nitrophenyl 3I-thiolaminaritrioside [Hordeum vulgare],3WLH_A Crystal Structure Analysis of Plant Exohydrolase [Hordeum vulgare subsp. vulgare],3WLJ_A Crystal structure of barley beta-D-glucan glucohydrolase isoenzyme EXO1 in complex with 3-deoxy-glucose [Hordeum vulgare subsp. vulgare],3WLK_X Crystal structure of barley beta-D-glucan glucohydrolase isoenzyme EXO1 in complex with 4-deoxy-glucose [Hordeum vulgare subsp. vulgare],3WLL_A Crystal structure of barley beta-D-glucan glucohydrolase isoenzyme EXO1 in complex with PEG400 [Hordeum vulgare subsp. vulgare],3WLM_A Crystal structure of barley beta-D-glucan glucohydrolase isoenzyme exo1 in complex with octyl-O-glucoside [Hordeum vulgare subsp. vulgare],3WLN_A Crystal structure of barley beta-D-glucan glucohydrolase isoenzyme EXO1 in complex with octyl-S-glucoside [Hordeum vulgare subsp. vulgare],3WLP_A Crystal Structure Analysis of Plant Exohydrolase [Hordeum vulgare subsp. vulgare] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| A7LXU3 | 3.95e-76 | 44 | 656 | 14 | 659 | Beta-glucosidase BoGH3B OS=Bacteroides ovatus (strain ATCC 8483 / DSM 1896 / JCM 5824 / BCRC 10623 / CCUG 4943 / NCTC 11153) OX=411476 GN=BACOVA_02659 PE=1 SV=1 |
| Q23892 | 6.38e-67 | 64 | 656 | 75 | 699 | Lysosomal beta glucosidase OS=Dictyostelium discoideum OX=44689 GN=gluA PE=1 SV=2 |
| Q56078 | 6.17e-58 | 61 | 656 | 26 | 648 | Periplasmic beta-glucosidase OS=Salmonella typhimurium (strain LT2 / SGSC1412 / ATCC 700720) OX=99287 GN=bglX PE=3 SV=2 |
| P33363 | 1.14e-57 | 64 | 656 | 33 | 648 | Periplasmic beta-glucosidase OS=Escherichia coli (strain K12) OX=83333 GN=bglX PE=3 SV=2 |
| T2KMH0 | 3.68e-45 | 61 | 656 | 29 | 604 | Beta-xylosidase OS=Formosa agariphila (strain DSM 15362 / KCTC 12365 / LMG 23005 / KMM 3901 / M-2Alg 35-1) OX=1347342 GN=BN863_22130 PE=1 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as SP
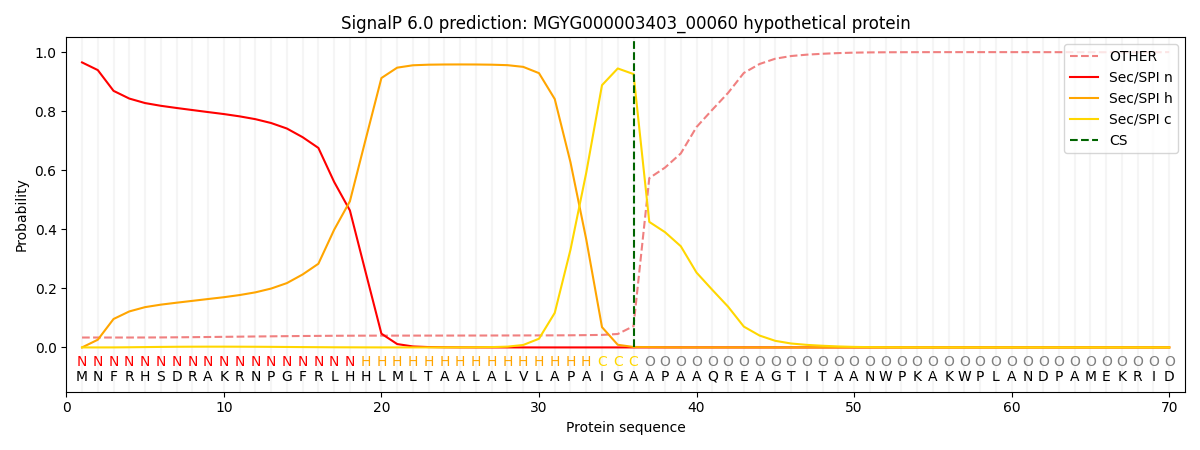
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 0.035997 | 0.961153 | 0.000879 | 0.001424 | 0.000291 | 0.000237 |