You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000003412_01721
You are here: Home > Sequence: MGYG000003412_01721
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | HGM12789 sp900766585 | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Firmicutes_A; Clostridia; Oscillospirales; Acutalibacteraceae; HGM12789; HGM12789 sp900766585 | |||||||||||
| CAZyme ID | MGYG000003412_01721 | |||||||||||
| CAZy Family | GH3 | |||||||||||
| CAZyme Description | Beta-hexosaminidase | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 103094; End: 105187 Strand: - | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH3 | 33 | 272 | 1.5e-72 | 0.9768518518518519 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| PLN03080 | PLN03080 | 2.14e-149 | 8 | 673 | 51 | 766 | Probable beta-xylosidase; Provisional |
| PRK15098 | PRK15098 | 2.33e-115 | 35 | 667 | 101 | 743 | beta-glucosidase BglX. |
| COG1472 | BglX | 1.23e-77 | 6 | 370 | 26 | 366 | Periplasmic beta-glucosidase and related glycosidases [Carbohydrate transport and metabolism]. |
| pfam01915 | Glyco_hydro_3_C | 1.07e-65 | 341 | 577 | 1 | 216 | Glycosyl hydrolase family 3 C-terminal domain. This domain is involved in catalysis and may be involved in binding beta-glucan. This domain is found associated with pfam00933. |
| pfam00933 | Glyco_hydro_3 | 2.72e-47 | 35 | 300 | 63 | 313 | Glycosyl hydrolase family 3 N terminal domain. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| AGC67275.1 | 1.57e-276 | 9 | 689 | 17 | 704 |
| ANW97778.1 | 1.57e-276 | 9 | 689 | 17 | 704 |
| ANX00304.1 | 1.57e-276 | 9 | 689 | 17 | 704 |
| AGI38341.1 | 1.57e-276 | 9 | 689 | 17 | 704 |
| AGX44636.1 | 1.04e-270 | 1 | 697 | 6 | 706 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 7VC7_A | 1.87e-104 | 9 | 580 | 25 | 600 | ChainA, xylan 1,4-beta-xylosidase [Phanerodontia chrysosporium] |
| 7VC6_A | 1.87e-104 | 9 | 580 | 25 | 600 | ChainA, xylan 1,4-beta-xylosidase [Phanerodontia chrysosporium] |
| 5Z87_A | 2.94e-101 | 35 | 684 | 120 | 776 | ChainA, EmGH1 [Aurantiacibacter marinus],5Z87_B Chain B, EmGH1 [Aurantiacibacter marinus] |
| 5A7M_A | 1.14e-86 | 9 | 580 | 52 | 623 | ChainA, BETA-XYLOSIDASE [Trichoderma reesei],5A7M_B Chain B, BETA-XYLOSIDASE [Trichoderma reesei] |
| 5AE6_A | 1.16e-86 | 9 | 580 | 52 | 623 | ChainA, BETA-XYLOSIDASE [Trichoderma reesei],5AE6_B Chain B, BETA-XYLOSIDASE [Trichoderma reesei] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| D5EY15 | 3.66e-148 | 6 | 676 | 31 | 847 | Xylan 1,4-beta-xylosidase OS=Prevotella ruminicola (strain ATCC 19189 / JCM 8958 / 23) OX=264731 GN=xyl3A PE=1 SV=1 |
| Q9SGZ5 | 2.05e-126 | 8 | 642 | 47 | 722 | Probable beta-D-xylosidase 7 OS=Arabidopsis thaliana OX=3702 GN=BXL7 PE=2 SV=2 |
| Q94KD8 | 1.76e-124 | 9 | 687 | 54 | 763 | Probable beta-D-xylosidase 2 OS=Arabidopsis thaliana OX=3702 GN=BXL2 PE=2 SV=1 |
| Q9FGY1 | 6.18e-123 | 7 | 674 | 57 | 760 | Beta-D-xylosidase 1 OS=Arabidopsis thaliana OX=3702 GN=BXL1 PE=1 SV=1 |
| A5JTQ2 | 2.00e-120 | 13 | 674 | 68 | 762 | Beta-xylosidase/alpha-L-arabinofuranosidase 1 (Fragment) OS=Medicago sativa subsp. varia OX=36902 GN=Xyl1 PE=1 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as OTHER
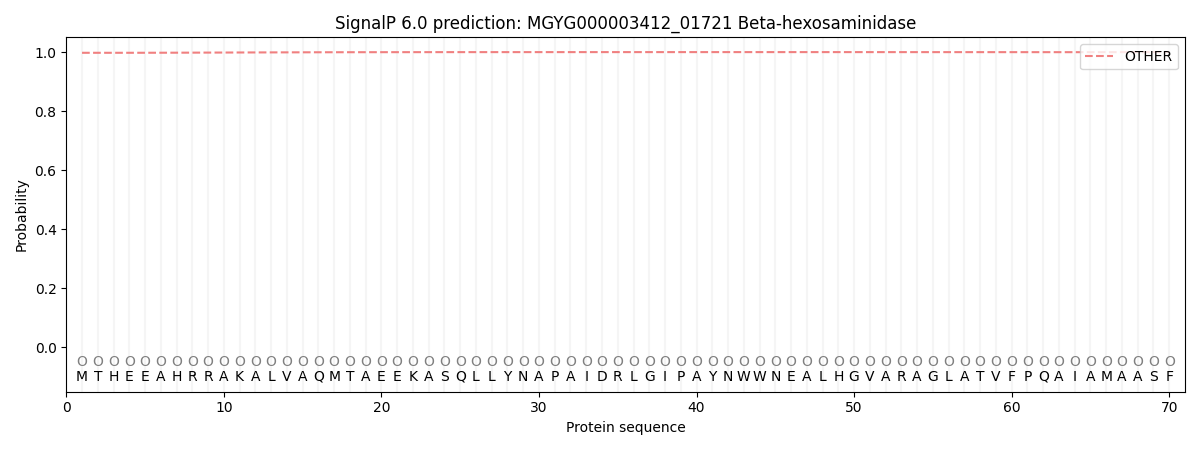
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 0.997889 | 0.001975 | 0.000050 | 0.000032 | 0.000012 | 0.000023 |
