You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000003430_00819
You are here: Home > Sequence: MGYG000003430_00819
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Eubacterium_R sp900766895 | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Firmicutes_A; Clostridia; Oscillospirales; Acutalibacteraceae; Eubacterium_R; Eubacterium_R sp900766895 | |||||||||||
| CAZyme ID | MGYG000003430_00819 | |||||||||||
| CAZy Family | GH28 | |||||||||||
| CAZyme Description | hypothetical protein | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 9200; End: 10744 Strand: + | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH28 | 84 | 449 | 2.1e-49 | 0.9107692307692308 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| COG5434 | Pgu1 | 1.17e-42 | 47 | 466 | 72 | 497 | Polygalacturonase [Carbohydrate transport and metabolism]. |
| pfam00295 | Glyco_hydro_28 | 7.97e-12 | 197 | 363 | 87 | 246 | Glycosyl hydrolases family 28. Glycosyl hydrolase family 28 includes polygalacturonase EC:3.2.1.15 as well as rhamnogalacturonase A(RGase A), EC:3.2.1.-. These enzymes are important in cell wall metabolism. |
| PLN03003 | PLN03003 | 1.68e-10 | 190 | 386 | 133 | 321 | Probable polygalacturonase At3g15720 |
| PLN02218 | PLN02218 | 7.18e-10 | 30 | 383 | 29 | 371 | polygalacturonase ADPG |
| PLN02188 | PLN02188 | 2.97e-07 | 58 | 363 | 37 | 316 | polygalacturonase/glycoside hydrolase family protein |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| AOZ96985.1 | 2.20e-83 | 31 | 487 | 12 | 508 |
| QGK73231.1 | 9.97e-40 | 52 | 513 | 18 | 460 |
| QDW21015.1 | 3.57e-39 | 52 | 513 | 18 | 460 |
| AOC96522.1 | 3.57e-39 | 52 | 513 | 18 | 460 |
| ADY37829.1 | 2.54e-38 | 59 | 513 | 26 | 471 |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| Q02096 | 2.16e-07 | 60 | 363 | 87 | 365 | Polygalacturonase OS=Persea americana OX=3435 PE=2 SV=1 |
| P18192 | 1.36e-06 | 244 | 363 | 226 | 336 | Endo-polygalacturonase OS=Pectobacterium carotovorum subsp. carotovorum OX=555 GN=peh PE=3 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as OTHER
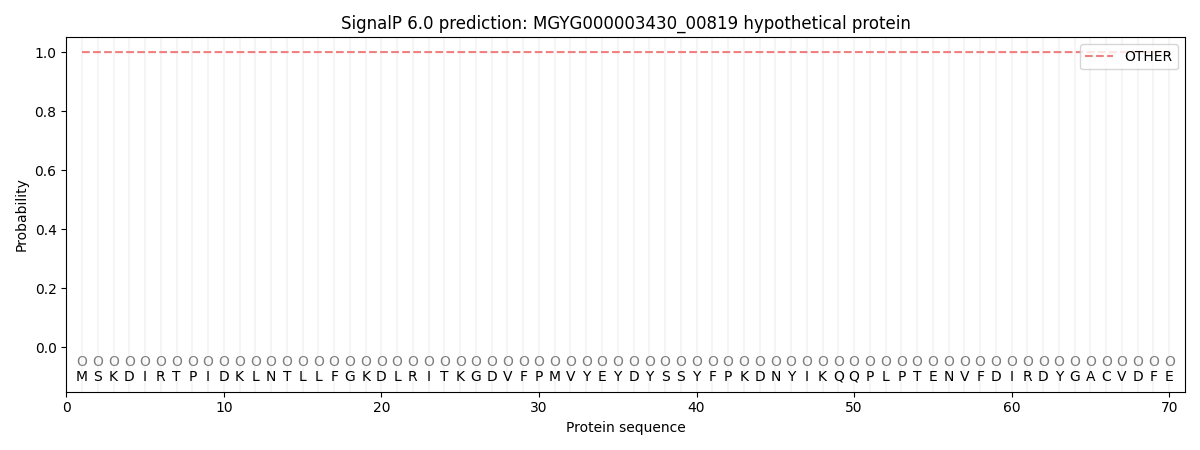
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 1.000068 | 0.000000 | 0.000000 | 0.000000 | 0.000000 | 0.000000 |
