You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000003445_00980
You are here: Home > Sequence: MGYG000003445_00980
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | RC9 sp900767375 | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Bacteroidota; Bacteroidia; Bacteroidales; UBA932; RC9; RC9 sp900767375 | |||||||||||
| CAZyme ID | MGYG000003445_00980 | |||||||||||
| CAZy Family | GH5 | |||||||||||
| CAZyme Description | hypothetical protein | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 8752; End: 10302 Strand: - | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH5 | 186 | 484 | 3.6e-98 | 0.9855072463768116 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| pfam00150 | Cellulase | 1.49e-50 | 186 | 483 | 15 | 266 | Cellulase (glycosyl hydrolase family 5). |
| COG2730 | BglC | 6.81e-22 | 150 | 483 | 48 | 357 | Aryl-phospho-beta-D-glucosidase BglC, GH1 family [Carbohydrate transport and metabolism]. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| ACA61144.1 | 9.99e-217 | 2 | 515 | 6 | 512 |
| ACA61160.1 | 1.34e-213 | 5 | 516 | 10 | 518 |
| AIT97140.1 | 2.53e-211 | 1 | 513 | 8 | 506 |
| ACA61149.1 | 4.31e-160 | 4 | 510 | 12 | 519 |
| ADX05703.1 | 1.78e-144 | 6 | 510 | 10 | 513 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 5D9N_A | 6.18e-92 | 144 | 512 | 11 | 353 | Crystalstructure of PbGH5A, a glycoside hydrolase family 5 member from Prevotella bryantii B14, in complex with the xyloglucan heptasaccharide XXXG [Prevotella bryantii],5D9N_B Crystal structure of PbGH5A, a glycoside hydrolase family 5 member from Prevotella bryantii B14, in complex with the xyloglucan heptasaccharide XXXG [Prevotella bryantii],5D9P_A Crystal structure of PbGH5A, a glycoside hydrolase family 5 enzyme from Prevotella bryantii B14, in complex with an inhibitory N-bromoacetylglycosylamine derivative of XXXG [Prevotella bryantii],5D9P_B Crystal structure of PbGH5A, a glycoside hydrolase family 5 enzyme from Prevotella bryantii B14, in complex with an inhibitory N-bromoacetylglycosylamine derivative of XXXG [Prevotella bryantii] |
| 5D9M_A | 4.88e-91 | 144 | 512 | 11 | 353 | Crystalstructure of PbGH5A, a glycoside hydrolase family 5 enzyme from Prevotella bryantii B14, E280A mutant in complex with the xyloglucan tetradecasaccharide XXXGXXXG [Prevotella bryantii],5D9M_B Crystal structure of PbGH5A, a glycoside hydrolase family 5 enzyme from Prevotella bryantii B14, E280A mutant in complex with the xyloglucan tetradecasaccharide XXXGXXXG [Prevotella bryantii],5D9O_A Crystal structure of PbGH5A, a glycoside hydrolase family 5 enzyme from Prevotella bryantii B14, E280A mutant in complex with cellotetraose [Prevotella bryantii],5D9O_B Crystal structure of PbGH5A, a glycoside hydrolase family 5 enzyme from Prevotella bryantii B14, E280A mutant in complex with cellotetraose [Prevotella bryantii] |
| 6D2W_A | 3.27e-87 | 144 | 512 | 434 | 776 | Crystalstructure of Prevotella bryantii endo-beta-mannanase/endo-beta-glucanase PbGH26A-GH5A [Prevotella bryantii B14],6D2W_B Crystal structure of Prevotella bryantii endo-beta-mannanase/endo-beta-glucanase PbGH26A-GH5A [Prevotella bryantii B14] |
| 3VDH_A | 5.27e-87 | 144 | 512 | 11 | 353 | Crystalstructure of PbGH5A, a glycoside hydrolase family 5 enzyme from Prevotella bryantii B14 [Prevotella bryantii],3VDH_B Crystal structure of PbGH5A, a glycoside hydrolase family 5 enzyme from Prevotella bryantii B14 [Prevotella bryantii] |
| 6XSO_A | 1.48e-80 | 141 | 510 | 17 | 376 | GH5-4broad specificity endoglucanase from an uncultured bacterium [uncultured bacterium],6XSO_B GH5-4 broad specificity endoglucanase from an uncultured bacterium [uncultured bacterium] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| Q12647 | 1.72e-60 | 150 | 483 | 29 | 328 | Endoglucanase B OS=Neocallimastix patriciarum OX=4758 GN=CELB PE=2 SV=1 |
| P17901 | 9.58e-60 | 148 | 483 | 49 | 365 | Endoglucanase A OS=Ruminiclostridium cellulolyticum (strain ATCC 35319 / DSM 5812 / JCM 6584 / H10) OX=394503 GN=celCCA PE=1 SV=1 |
| P23661 | 2.04e-59 | 146 | 513 | 71 | 402 | Endoglucanase B OS=Ruminococcus albus OX=1264 GN=celB PE=3 SV=1 |
| P16216 | 7.22e-59 | 146 | 483 | 69 | 363 | Endoglucanase 1 OS=Ruminococcus albus OX=1264 GN=Eg I PE=1 SV=1 |
| P54937 | 1.78e-58 | 146 | 512 | 39 | 380 | Endoglucanase A OS=Clostridium longisporum OX=1523 GN=celA PE=1 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as LIPO
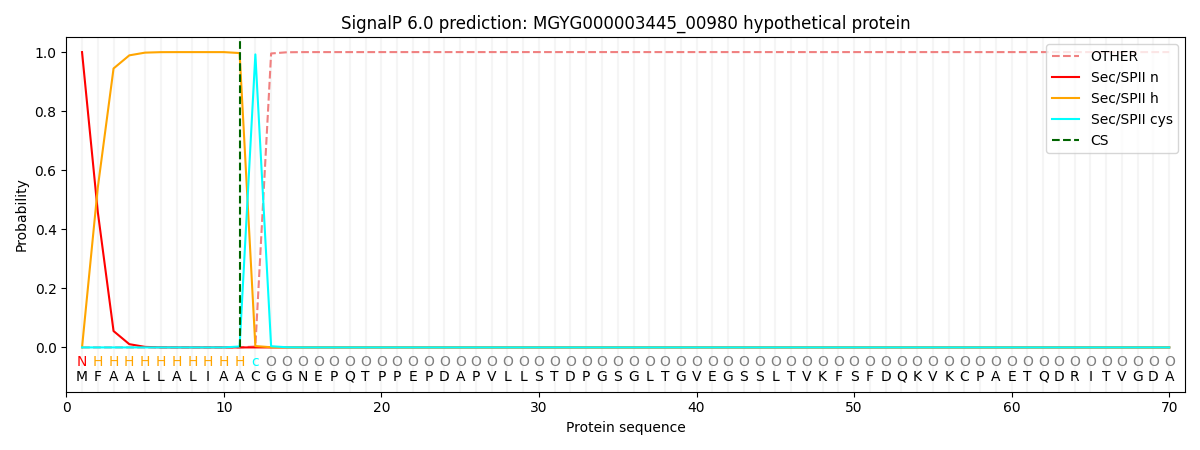
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 0.000000 | 0.000001 | 1.000027 | 0.000000 | 0.000000 | 0.000000 |
