You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000003512_00294
You are here: Home > Sequence: MGYG000003512_00294
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | UBA11524 sp900769075 | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Firmicutes_A; Clostridia_A; Christensenellales; CAG-74; UBA11524; UBA11524 sp900769075 | |||||||||||
| CAZyme ID | MGYG000003512_00294 | |||||||||||
| CAZy Family | GH9 | |||||||||||
| CAZyme Description | Cellodextrinase | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 36940; End: 38472 Strand: - | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH9 | 88 | 506 | 8e-104 | 0.9952153110047847 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| pfam00759 | Glyco_hydro_9 | 2.24e-91 | 98 | 505 | 9 | 374 | Glycosyl hydrolase family 9. |
| cd02850 | E_set_Cellulase_N | 7.78e-21 | 2 | 83 | 1 | 86 | N-terminal Early set domain associated with the catalytic domain of cellulase. E or "early" set domains are associated with the catalytic domain of cellulases at the N-terminal end. Cellulases are O-glycosyl hydrolases (GHs) that hydrolyze beta 1-4 glucosidic bonds in cellulose. They are usually categorized into either exoglucanases, which sequentially release terminal sugar units from the cellulose chain, or endoglucanases, which also attack the chain internally. The N-terminal domain of cellulase may be related to the immunoglobulin and/or fibronectin type III superfamilies. These domains are associated with different types of catalytic domains at either the N-terminal or C-terminal end and may be involved in homodimeric/tetrameric/dodecameric interactions. Members of this family include members of the alpha amylase family, sialidase, galactose oxidase, cellulase, cellulose, hyaluronate lyase, chitobiase, and chitinase, among others. |
| PLN02613 | PLN02613 | 5.24e-16 | 128 | 445 | 63 | 410 | endoglucanase |
| pfam02927 | CelD_N | 9.43e-15 | 2 | 78 | 2 | 83 | Cellulase N-terminal ig-like domain. |
| PLN02420 | PLN02420 | 7.20e-12 | 120 | 508 | 72 | 506 | endoglucanase |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| AEJ61964.1 | 1.40e-133 | 3 | 510 | 210 | 734 |
| ADN02583.1 | 2.31e-131 | 3 | 510 | 208 | 732 |
| CBL16782.1 | 4.18e-130 | 6 | 506 | 204 | 716 |
| ADH59533.1 | 6.91e-130 | 3 | 506 | 9 | 536 |
| QUL57275.1 | 1.11e-125 | 3 | 506 | 9 | 537 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 5E2J_A | 2.03e-110 | 6 | 508 | 35 | 554 | Crystalstructure of single mutant thermostable endoglucanase (D468A) from Alicyclobacillus acidocaldarius [Alicyclobacillus acidocaldarius subsp. acidocaldarius],5E2J_B Crystal structure of single mutant thermostable endoglucanase (D468A) from Alicyclobacillus acidocaldarius [Alicyclobacillus acidocaldarius subsp. acidocaldarius] |
| 3EZ8_A | 2.20e-110 | 6 | 508 | 12 | 531 | CrystalStructure of endoglucanase Cel9A from the thermoacidophilic Alicyclobacillus acidocaldarius [Alicyclobacillus acidocaldarius subsp. acidocaldarius],3GZK_A Structure of A. Acidocaldarius Cellulase CelA [Alicyclobacillus acidocaldarius subsp. acidocaldarius],3H2W_A Structure of A. acidocaldarius cellulase CelA in complex with cellobiose [Alicyclobacillus acidocaldarius subsp. acidocaldarius],3H3K_A Structure of A. acidocaldarius cellulase CelA in complex with cellotetraose [Alicyclobacillus acidocaldarius subsp. acidocaldarius],3RX5_A structure of AaCel9A in complex with cellotriose-like isofagomine [Alicyclobacillus acidocaldarius subsp. acidocaldarius],3RX7_A Structure of AaCel9A in complex with cellotetraose-like isofagomine [Alicyclobacillus acidocaldarius subsp. acidocaldarius],3RX8_A structure of AaCel9A in complex with cellobiose-like isofagomine [Alicyclobacillus acidocaldarius subsp. acidocaldarius] |
| 3X17_A | 7.43e-85 | 3 | 506 | 18 | 554 | Crystalstructure of metagenome-derived glycoside hydrolase family 9 endoglucanase [uncultured bacterium],3X17_B Crystal structure of metagenome-derived glycoside hydrolase family 9 endoglucanase [uncultured bacterium] |
| 4CJ0_A | 6.94e-71 | 3 | 510 | 30 | 549 | ChainA, ENDOGLUCANASE D [Acetivibrio thermocellus],4CJ1_A Chain A, ENDOGLUCANASE D [Acetivibrio thermocellus] |
| 1CLC_A | 9.20e-71 | 3 | 510 | 44 | 563 | ChainA, ENDOGLUCANASE CELD; EC: 3.2.1.4 [Acetivibrio thermocellus] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| P23658 | 1.82e-86 | 5 | 508 | 6 | 545 | Cellodextrinase OS=Butyrivibrio fibrisolvens OX=831 GN=ced1 PE=1 SV=1 |
| P0C2S4 | 3.80e-70 | 3 | 510 | 30 | 549 | Endoglucanase D (Fragment) OS=Acetivibrio thermocellus OX=1515 GN=celD PE=1 SV=1 |
| A3DDN1 | 6.13e-70 | 3 | 510 | 54 | 573 | Endoglucanase D OS=Acetivibrio thermocellus (strain ATCC 27405 / DSM 1237 / JCM 9322 / NBRC 103400 / NCIMB 10682 / NRRL B-4536 / VPI 7372) OX=203119 GN=celD PE=1 SV=1 |
| Q05156 | 1.65e-62 | 3 | 504 | 185 | 738 | Cellulase 1 OS=Streptomyces reticuli OX=1926 GN=cel1 PE=1 SV=1 |
| A7LXT3 | 3.18e-58 | 3 | 508 | 32 | 578 | Xyloglucan-specific endo-beta-1,4-glucanase BoGH9A OS=Bacteroides ovatus (strain ATCC 8483 / DSM 1896 / JCM 5824 / BCRC 10623 / CCUG 4943 / NCTC 11153) OX=411476 GN=BACOVA_02649 PE=1 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as OTHER
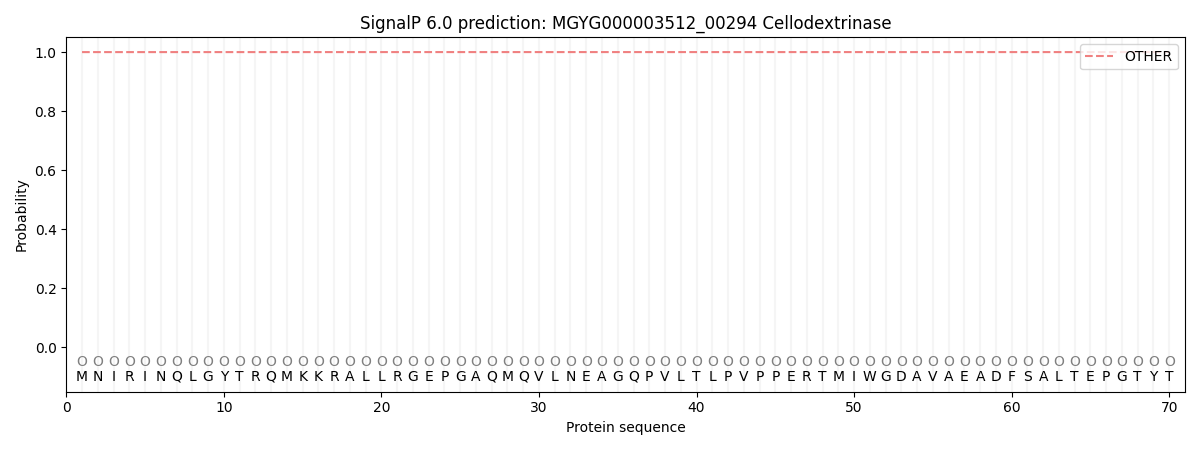
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 1.000044 | 0.000000 | 0.000000 | 0.000000 | 0.000000 | 0.000000 |
