You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000003520_01864
You are here: Home > Sequence: MGYG000003520_01864
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | RC9 sp900769145 | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Bacteroidota; Bacteroidia; Bacteroidales; UBA932; RC9; RC9 sp900769145 | |||||||||||
| CAZyme ID | MGYG000003520_01864 | |||||||||||
| CAZy Family | GH5 | |||||||||||
| CAZyme Description | hypothetical protein | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 30040; End: 31284 Strand: + | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH5 | 91 | 342 | 2.8e-73 | 0.9883268482490273 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| pfam00150 | Cellulase | 3.01e-30 | 92 | 336 | 15 | 266 | Cellulase (glycosyl hydrolase family 5). |
| COG2730 | BglC | 1.53e-21 | 51 | 280 | 16 | 250 | Aryl-phospho-beta-D-glucosidase BglC, GH1 family [Carbohydrate transport and metabolism]. |
| pfam13204 | DUF4038 | 3.74e-04 | 129 | 255 | 69 | 176 | Protein of unknown function (DUF4038). A family of putative cellulases. |
| pfam01261 | AP_endonuc_2 | 0.006 | 141 | 254 | 63 | 152 | Xylose isomerase-like TIM barrel. This TIM alpha/beta barrel structure is found in xylose isomerase and in endonuclease IV (EC:3.1.21.2). This domain is also found in the N termini of bacterial myo-inositol catabolism proteins. These are involved in the myo-inositol catabolism pathway, and is required for growth on myo-inositol in Rhizobium leguminosarum bv. viciae. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| QUT89172.1 | 6.32e-109 | 3 | 414 | 2 | 415 |
| BBE18442.1 | 1.97e-108 | 7 | 414 | 8 | 417 |
| ALJ59797.1 | 1.44e-107 | 3 | 414 | 2 | 415 |
| QDO68125.1 | 1.44e-107 | 3 | 414 | 2 | 415 |
| ADV42635.1 | 1.39e-105 | 5 | 414 | 2 | 416 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 1CEC_A | 9.04e-15 | 97 | 262 | 22 | 176 | ChainA, ENDOGLUCANASE CELC [Acetivibrio thermocellus] |
| 1CEN_A | 2.20e-14 | 97 | 262 | 22 | 176 | ChainA, CELLULASE CELC [Acetivibrio thermocellus],1CEO_A Chain A, CELLULASE CELC [Acetivibrio thermocellus] |
| 3AMC_A | 6.49e-12 | 104 | 338 | 34 | 288 | Crystalstructures of Thermotoga maritima Cel5A, apo form and dimer/au [Thermotoga maritima MSB8],3AMC_B Crystal structures of Thermotoga maritima Cel5A, apo form and dimer/au [Thermotoga maritima MSB8],3AMD_A Crystal structures of Thermotoga maritima Cel5A, apo form and tetramer/au [Thermotoga maritima MSB8],3AMD_B Crystal structures of Thermotoga maritima Cel5A, apo form and tetramer/au [Thermotoga maritima MSB8],3AMD_C Crystal structures of Thermotoga maritima Cel5A, apo form and tetramer/au [Thermotoga maritima MSB8],3AMD_D Crystal structures of Thermotoga maritima Cel5A, apo form and tetramer/au [Thermotoga maritima MSB8],3MMU_A Crystal structure of endoglucanase Cel5A from the hyperthermophilic Thermotoga maritima [Thermotoga maritima],3MMU_B Crystal structure of endoglucanase Cel5A from the hyperthermophilic Thermotoga maritima [Thermotoga maritima],3MMU_C Crystal structure of endoglucanase Cel5A from the hyperthermophilic Thermotoga maritima [Thermotoga maritima],3MMU_D Crystal structure of endoglucanase Cel5A from the hyperthermophilic Thermotoga maritima [Thermotoga maritima],3MMU_E Crystal structure of endoglucanase Cel5A from the hyperthermophilic Thermotoga maritima [Thermotoga maritima],3MMU_F Crystal structure of endoglucanase Cel5A from the hyperthermophilic Thermotoga maritima [Thermotoga maritima],3MMU_G Crystal structure of endoglucanase Cel5A from the hyperthermophilic Thermotoga maritima [Thermotoga maritima],3MMU_H Crystal structure of endoglucanase Cel5A from the hyperthermophilic Thermotoga maritima [Thermotoga maritima],3MMW_A Crystal structure of endoglucanase Cel5A from the hyperthermophilic Thermotoga maritima [Thermotoga maritima],3MMW_B Crystal structure of endoglucanase Cel5A from the hyperthermophilic Thermotoga maritima [Thermotoga maritima],3MMW_C Crystal structure of endoglucanase Cel5A from the hyperthermophilic Thermotoga maritima [Thermotoga maritima],3MMW_D Crystal structure of endoglucanase Cel5A from the hyperthermophilic Thermotoga maritima [Thermotoga maritima] |
| 6ZB9_A | 2.35e-11 | 53 | 285 | 3 | 254 | ChainA, Exo-beta-1,3-glucanase [uncultured bacterium],6ZB9_B Chain B, Exo-beta-1,3-glucanase [uncultured bacterium] |
| 3AZR_A | 3.77e-11 | 104 | 338 | 34 | 288 | DiverseSubstrates Recognition Mechanism Revealed by Thermotoga maritima Cel5A Structures in Complex with Cellobiose [Thermotoga maritima MSB8],3AZR_B Diverse Substrates Recognition Mechanism Revealed by Thermotoga maritima Cel5A Structures in Complex with Cellobiose [Thermotoga maritima MSB8],3AZS_A Diverse Substrates Recognition Mechanism Revealed by Thermotoga maritima Cel5A Structures in Complex with Mannotriose [Thermotoga maritima MSB8],3AZS_B Diverse Substrates Recognition Mechanism Revealed by Thermotoga maritima Cel5A Structures in Complex with Mannotriose [Thermotoga maritima MSB8],3AZT_A Diverse Substrates Recognition Mechanism Revealed by Thermotoga maritima Cel5A Structures in Complex with Cellotetraose [Thermotoga maritima MSB8],3AZT_B Diverse Substrates Recognition Mechanism Revealed by Thermotoga maritima Cel5A Structures in Complex with Cellotetraose [Thermotoga maritima MSB8],3AZT_C Diverse Substrates Recognition Mechanism Revealed by Thermotoga maritima Cel5A Structures in Complex with Cellotetraose [Thermotoga maritima MSB8],3AZT_D Diverse Substrates Recognition Mechanism Revealed by Thermotoga maritima Cel5A Structures in Complex with Cellotetraose [Thermotoga maritima MSB8] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| W8QRE4 | 1.90e-35 | 35 | 267 | 3 | 229 | Beta-xylosidase OS=Phanerodontia chrysosporium OX=2822231 GN=Xyl5 PE=1 SV=2 |
| Q8NKF9 | 1.88e-14 | 53 | 356 | 37 | 361 | Glucan 1,3-beta-glucosidase OS=Candida oleophila OX=45573 GN=EXG1 PE=3 SV=1 |
| A3DJ77 | 2.03e-14 | 97 | 262 | 22 | 176 | Endoglucanase C OS=Acetivibrio thermocellus (strain ATCC 27405 / DSM 1237 / JCM 9322 / NBRC 103400 / NCIMB 10682 / NRRL B-4536 / VPI 7372) OX=203119 GN=celC PE=3 SV=1 |
| P23340 | 2.03e-14 | 97 | 262 | 22 | 176 | Endoglucanase C307 OS=Clostridium sp. (strain F1) OX=1508 GN=celC307 PE=1 SV=1 |
| P0C2S3 | 2.74e-14 | 97 | 262 | 22 | 176 | Endoglucanase C OS=Acetivibrio thermocellus OX=1515 GN=celC PE=1 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as LIPO
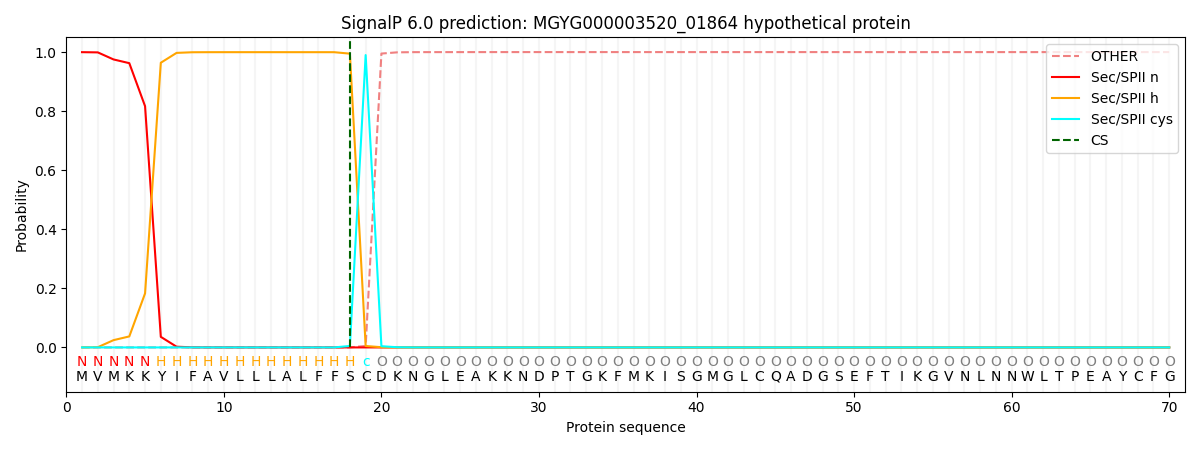
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 0.000000 | 0.000235 | 0.999791 | 0.000000 | 0.000000 | 0.000000 |
