You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000003534_00408
You are here: Home > Sequence: MGYG000003534_00408
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Treponema_D sp900769325 | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Spirochaetota; Spirochaetia; Treponematales; Treponemataceae; Treponema_D; Treponema_D sp900769325 | |||||||||||
| CAZyme ID | MGYG000003534_00408 | |||||||||||
| CAZy Family | GH57 | |||||||||||
| CAZyme Description | hypothetical protein | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 7105; End: 9099 Strand: + | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH57 | 23 | 367 | 1.5e-35 | 0.9295039164490861 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| cd10793 | GH57N_TLGT_like | 5.30e-30 | 22 | 279 | 18 | 277 | N-terminal catalytic domain of 4-alpha-glucanotransferase; glycoside hydrolase family 57 (GH57). 4-alpha-glucanotransferase (TLGT, EC 2.4.1.25) plays a key role in the maltose metabolism. It catalyzes the disproportionation of amylose and the formation of large cyclic alpha-1,4-glucan (cycloamylose) from linear amylose. TLGT functions as a homodimer. Each monomer is composed of two domains, an N-terminal catalytic domain with a (beta/alpha)7 barrel fold and a C-terminal domain with a twisted beta-sandwich fold. Some family members have been designated as alpha-amylases, such as the heat-stable eubacterial amylase from Dictyoglomus thermophilum (DtAmyA) and the extremely thermostable archaeal amylase from Pyrococcus furiosus(PfAmyA). However, both of these proteins are 4-alpha-glucanotransferases. DtAmyA was shown to have transglycosylating activity and PfAmyA exhibits 4-alpha-glucanotransferase activity. |
| pfam03065 | Glyco_hydro_57 | 1.14e-16 | 26 | 268 | 51 | 293 | Glycosyl hydrolase family 57. This family includes alpha-amylase (EC:3.2.1.1), 4--glucanotransferase (EC:2.4.1.-) and amylopullulanase enzymes. |
| pfam09095 | DUF1926 | 3.24e-07 | 388 | 542 | 1 | 188 | Domain of unknown function (DUF1926). Members of this family, which are found in a set of prokaryotic transferases, adopt a beta-sandwich fold, in which two layers of anti-parallel beta-sheets are arranged in a nearly parallel fashion. The exact function of this family is, as yet, unknown, however it has been proposed that they may play a role in transglycosylation reactions. |
| cd01022 | GH57N_like | 3.78e-06 | 36 | 272 | 41 | 301 | N-terminal catalytic domain of heat stable retaining glycoside hydrolase family 57. Glycoside hydrolase family 57(GH57) is a chiefly prokaryotic family with the majority of thermostable enzymes coming from extremophiles (many of these are archaeal hyperthermophiles), which exhibit the enzyme specificities of alpha-amylase (EC 3.2.1.1), 4-alpha-glucanotransferase (EC 2.4.1.25), amylopullulanase (EC 3.2.1.1/41), and alpha-galactosidase (EC 3.2.1.22). This family also includes many hypothetical proteins with uncharacterized activity and specificity. GH57s cleave alpha-glycosidic bonds by employing a retaining mechanism, which involves a glycosyl-enzyme intermediate, allowing transglycosylation. |
| cd10798 | GH57N_like_1 | 6.20e-05 | 46 | 153 | 65 | 190 | Uncharacterized subfamily of glycoside hydrolase family 57 (GH57). This subfamily of uncharacterized bacterial proteins, shows high sequence homology to glycoside hydrolase family 57 (GH57). Glycoside hydrolase family 57(GH57) is a chiefly prokaryotic family with the majority of thermostable enzymes coming from extremophiles (many of these are archaeal hyperthermophiles), which exhibit the enzyme specificities of alpha-amylase (EC 3.2.1.1), 4-alpha-glucanotransferase (EC 2.4.1.25), amylopullulanase (EC 3.2.1.1/41), and alpha-galactosidase (EC 3.2.1.22). |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| QNL97410.1 | 5.06e-205 | 19 | 664 | 18 | 665 |
| QTQ16225.1 | 5.24e-188 | 26 | 663 | 26 | 656 |
| QTQ11807.1 | 7.41e-188 | 26 | 663 | 26 | 656 |
| QSI02287.1 | 1.45e-186 | 1 | 658 | 1 | 656 |
| QTQ14015.1 | 1.91e-185 | 26 | 663 | 26 | 656 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 1K1X_A | 6.95e-27 | 1 | 433 | 1 | 438 | Crystalstructure of 4-alpha-glucanotransferase from thermococcus litoralis [Thermococcus litoralis],1K1X_B Crystal structure of 4-alpha-glucanotransferase from thermococcus litoralis [Thermococcus litoralis],1K1Y_A Crystal structure of thermococcus litoralis 4-alpha-glucanotransferase complexed with acarbose [Thermococcus litoralis],1K1Y_B Crystal structure of thermococcus litoralis 4-alpha-glucanotransferase complexed with acarbose [Thermococcus litoralis] |
| 1K1W_A | 1.94e-23 | 24 | 433 | 24 | 438 | Crystalstructure of 4-alpha-glucanotransferase from thermococcus litoralis [Thermococcus litoralis] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| P09961 | 4.44e-31 | 23 | 436 | 24 | 453 | Alpha-amylase 1 OS=Dictyoglomus thermophilum (strain ATCC 35947 / DSM 3960 / H-6-12) OX=309799 GN=amyA PE=1 SV=2 |
| P49067 | 2.76e-26 | 24 | 563 | 25 | 588 | Alpha-amylase OS=Pyrococcus furiosus (strain ATCC 43587 / DSM 3638 / JCM 8422 / Vc1) OX=186497 GN=amyA PE=1 SV=2 |
| O32462 | 3.80e-26 | 1 | 433 | 1 | 438 | 4-alpha-glucanotransferase OS=Thermococcus litoralis (strain ATCC 51850 / DSM 5473 / JCM 8560 / NS-C) OX=523849 GN=jgt PE=1 SV=1 |
| Q9V298 | 1.14e-24 | 23 | 563 | 23 | 596 | Alpha-amylase OS=Pyrococcus abyssi (strain GE5 / Orsay) OX=272844 GN=amyA PE=3 SV=1 |
| O57932 | 4.20e-23 | 23 | 473 | 23 | 492 | Alpha-amylase OS=Pyrococcus horikoshii (strain ATCC 700860 / DSM 12428 / JCM 9974 / NBRC 100139 / OT-3) OX=70601 GN=amyA PE=3 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as OTHER
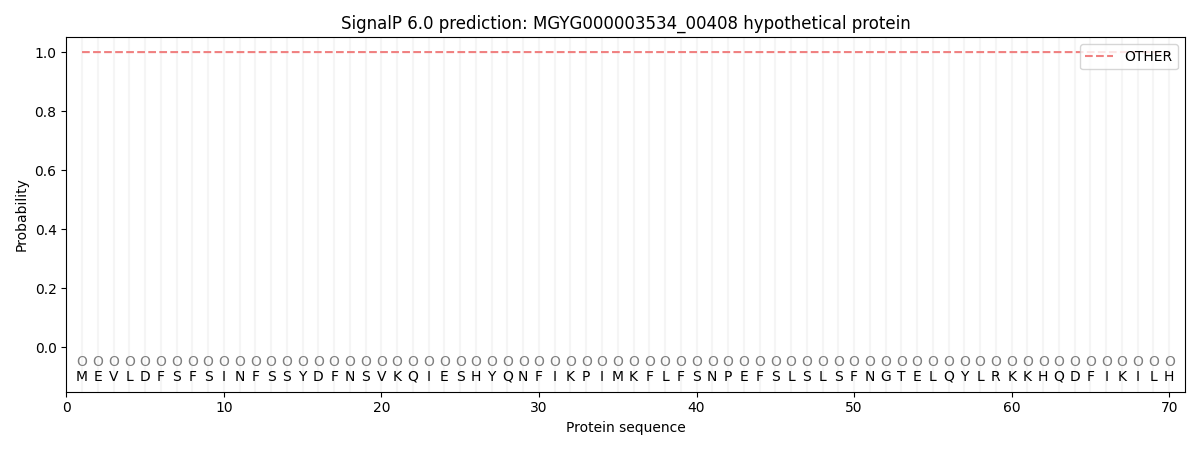
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 1.000052 | 0.000000 | 0.000000 | 0.000000 | 0.000000 | 0.000000 |
